skull_CTA
Apply a specialized skull-stripping pipeline to a single angiography volume using intensity preprocessing, Gaussian smoothing, and FSL BET for brain extraction.
skull_CTA(
nii_path: str,
output_path: str,
f_value: float = 0.1,
clip_value: tuple = (0, 200),
cleanup: bool = False,
debug: bool = False
) -> None
Overview
This function implements a multi-stage skull-stripping pipeline specifically designed for contrast-enhanced angiography imaging. The pipeline combines intensity preprocessing with FSL’s Brain Extraction Tool (BET) to isolate brain tissue and vasculature from skull and non-brain structures.
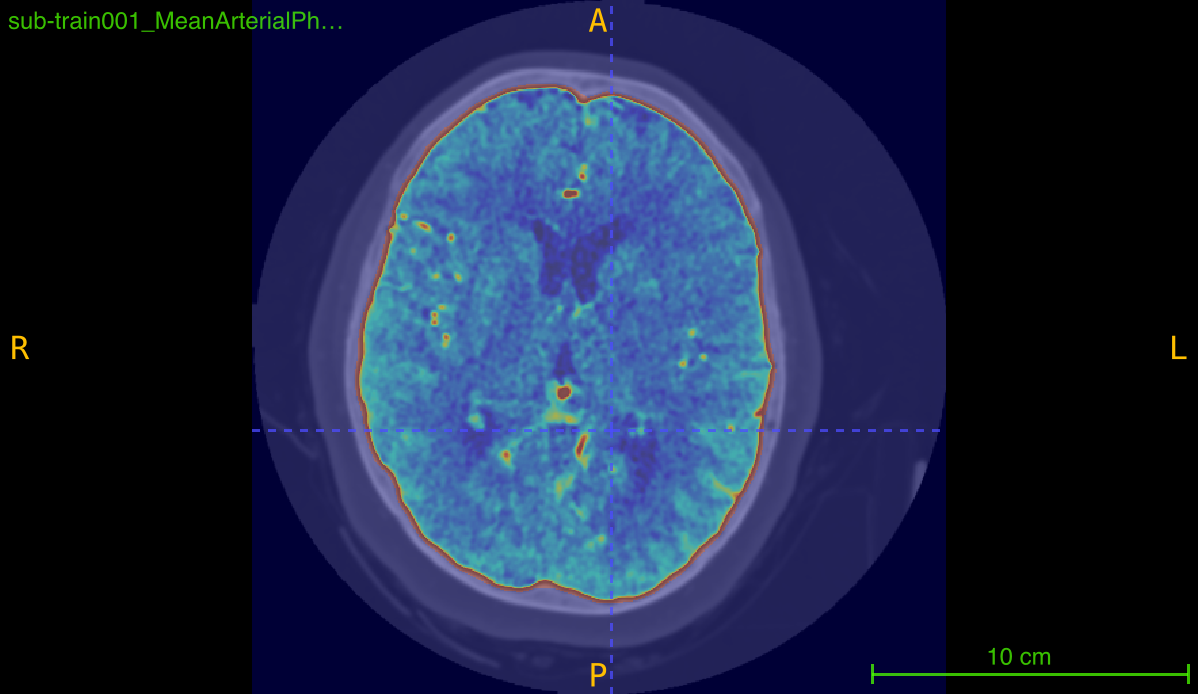
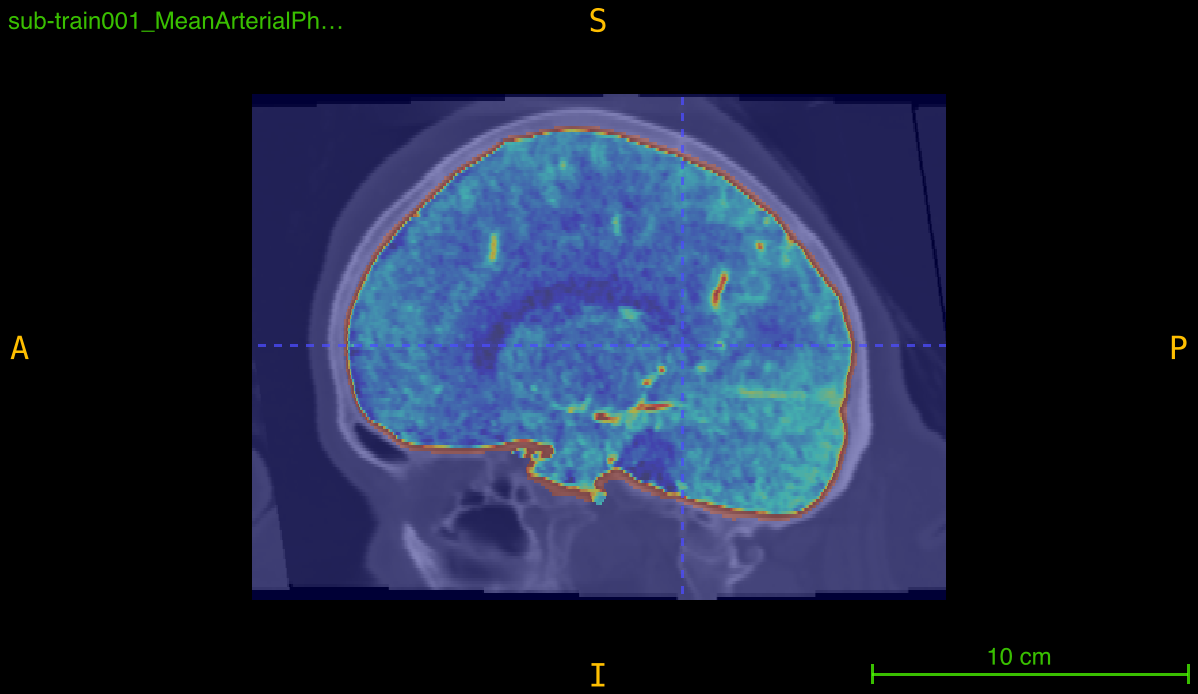
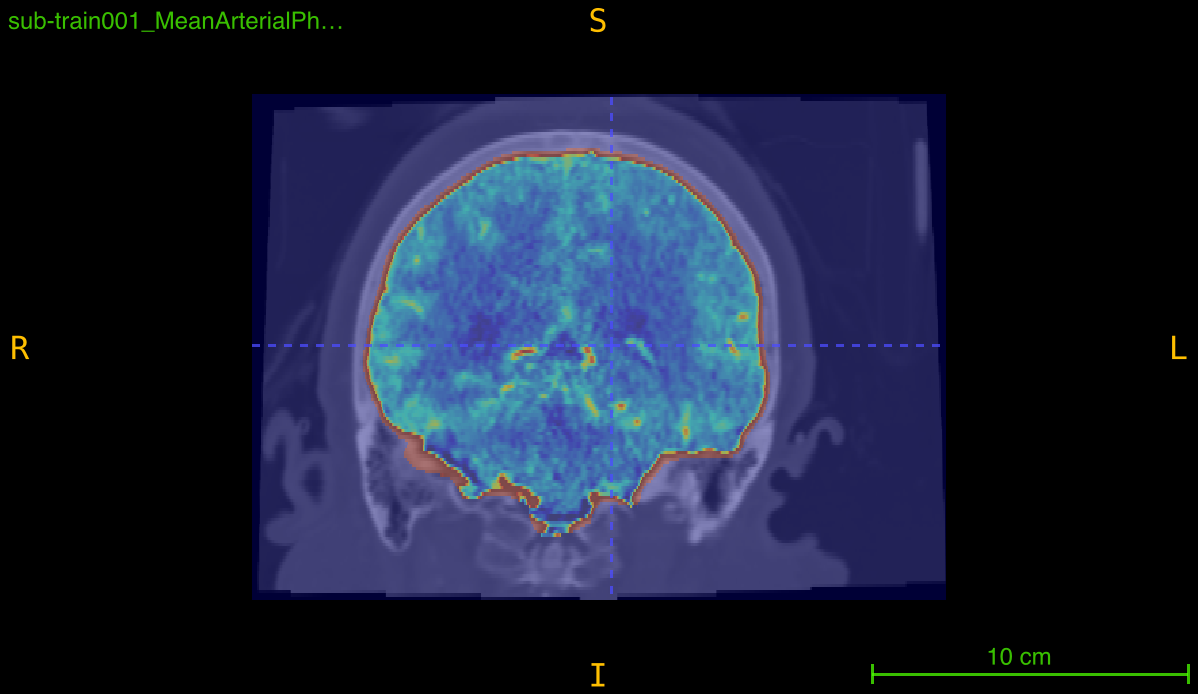
Processing pipeline:
- Initial thresholding: Clip intensities to [0, 100] range
- Gaussian smoothing: Reduce noise with sigma=1
- Secondary thresholding: Re-clip to [0, 100] after smoothing
- BET skull-stripping: Remove skull using FSL BET
- Final clipping: Normalize to desired intensity range
This is essential for:
- Isolating brain vasculature in angiography scans
- Removing skull interference in vessel detection
- Standardizing intensity ranges for analysis
- Preparing data for vessel segmentation
- Reducing false positives from bone in detection tasks
Parameters
| Name | Type | Default | Description |
|---|---|---|---|
nii_path | str | required | Path to the input angiography volume in .nii.gz format. |
output_path | str | required | Directory where all outputs will be saved. Created automatically if it doesn’t exist. |
f_value | float | 0.1 | Fractional intensity threshold for BET (0-1). Lower values = more aggressive stripping. |
clip_value | tuple | (0, 200) | Intensity range (min, max) for final clipping after skull removal. |
cleanup | bool | False | If True, deletes intermediate files. Mask and final clipped volume always retained. |
debug | bool | False | If True, prints detailed information about output file locations. |
Returns
None – The function saves processed volumes and mask to disk.
Output Files
The function generates multiple files during processing:
Intermediate Files (removed if cleanup=True)
| File | Description |
|---|---|
<PREFIX>_th.nii.gz | After initial thresholding [0-100] |
<PREFIX>_th_sm.nii.gz | After Gaussian smoothing (sigma=1) |
<PREFIX>_th_sm_th.nii.gz | After secondary thresholding [0-100] |
<PREFIX>_skulled.nii.gz | BET output before final clipping |
Final Output Files (always retained)
| File | Description |
|---|---|
<PREFIX>_skulled_mask.nii.gz | Binary brain mask from BET |
<PREFIX>_skulled_clipped.nii.gz | Final skull-stripped and intensity-clipped volume |
Example: Input scan_042.nii.gz produces:
scan_042_skulled_mask.nii.gzscan_042_skulled_clipped.nii.gz
Processing Pipeline Details
Step 1: Initial Thresholding
fslmaths input.nii.gz -thr 0 -uthr 100 output_th.nii.gz
Clips intensity values to [0, 100] range to remove extreme values and background.
Step 2: Gaussian Smoothing
fslmaths input_th.nii.gz -s 1 output_th_sm.nii.gz
Applies Gaussian blur with sigma=1 to reduce noise while preserving structures.
Step 3: Secondary Thresholding
fslmaths input_th_sm.nii.gz -thr 0 -uthr 100 output_th_sm_th.nii.gz
Re-applies [0, 100] clipping after smoothing to ensure intensity range.
Step 4: BET Skull-Stripping
bet input_th_sm_th.nii.gz output_skulled.nii.gz -R -f 0.1 -g 0 -m
Parameters:
-R: Robust brain center estimation-f 0.1: Fractional intensity threshold (default)-g 0: No vertical gradient in intensity-m: Generate binary mask
Step 5: Final Intensity Clipping
clipped = np.clip(skulled_data, min, max)
Normalizes intensity range to specified bounds (default [0, 200]).
BET f_value Parameter
The f_value parameter controls skull removal aggressiveness:
| f_value | Effect | When to Use |
|---|---|---|
| 0.05 | Very aggressive, removes more | Thick skull, heavy calcification |
| 0.1 | Aggressive (default) | Standard angiography scans |
| 0.2 | Moderate | Thin skull, pediatric cases |
| 0.3-0.5 | Conservative, preserves more | Ensure all brain tissue retained |
Guidelines:
- Lower (0.05-0.15): More skull removed, risk of brain removal
- Higher (0.2-0.5): Less aggressive, may leave some skull
- Start with 0.1 and adjust based on visual inspection
Intensity Clipping
The clip_value tuple defines the final intensity normalization:
| Range | Effect | Use Case |
|---|---|---|
| (0, 100) | Conservative, soft tissue focus | Basic tissue analysis |
| (0, 200) | Standard angiography range (default) | Most vessel analysis tasks |
| (0, 300) | Include high-intensity contrast | Bright vessel enhancement |
| (0, 500) | Very inclusive, may include artifacts | Preserve all contrast regions |
FSL Requirements
Critical: This function requires FSL (FMRIB Software Library) installation:
Installation
- Download from https://fsl.fmrib.ox.ac.uk/
- Follow platform-specific installation instructions
- Configure environment variables (typically in
.bashrcor.bash_profile)
Verification
# Check FSL installation
which fslmaths # Should return path to executable
which bet # Should return path to executable
# Test BET
bet # Should display help information
Environment Setup
FSL requires proper environment configuration:
# Typical FSL setup (add to .bashrc)
FSLDIR=/usr/local/fsl
. ${FSLDIR}/etc/fslconf/fsl.sh
PATH=${FSLDIR}/bin:${PATH}
export FSLDIR PATH
Execution Requirements
- Run from terminal: Execute Python scripts from command line, not IDE
- Environment variables: FSL paths must be in shell environment
- Command:
python3 script.py(not from IDE’s run button)
Exceptions
| Exception | Condition |
|---|---|
FileNotFoundError | The input file does not exist |
ValueError | File is not in .nii.gz format |
ValueError | Input data is not 3D |
RuntimeError | FSL command execution failed |
Usage Notes
- Input Format: Only
.nii.gzfiles are accepted - 3D Volumes Required: Input must be a 3D NIfTI image
- Pre-centered Volumes: Input should be centered on brain region
- No FOV Cropping: Original dimensions are preserved
- FSL Dependency: FSL must be installed and properly configured
- Output Directory: Automatically created if it doesn’t exist
- Intermediate Files: Can be removed with
cleanup=Trueto save space
Examples
Basic Usage
Process a single angiography volume:
from nidataset.preprocessing import skull_CTA
skull_CTA(
nii_path="angiography/patient_001.nii.gz",
output_path="processed/patient_001/",
f_value=0.1,
clip_value=(0, 200)
)
# Output: processed/patient_001/patient_001_skulled_clipped.nii.gz
# processed/patient_001/patient_001_skulled_mask.nii.gz
With Cleanup
Remove intermediate files to save disk space:
skull_CTA(
nii_path="data/scan.nii.gz",
output_path="data/processed/",
f_value=0.1,
clip_value=(0, 200),
cleanup=True, # Remove intermediate files
debug=True
)
# Prints: Skull-stripped image saved at: 'data/processed/scan_skulled_clipped.nii.gz'
# Skull mask saved at: 'data/processed/scan_skulled_mask.nii.gz'
Aggressive Skull Removal
Use lower f_value for thick skull:
skull_CTA(
nii_path="thick_skull/scan.nii.gz",
output_path="aggressive_strip/",
f_value=0.05, # More aggressive
clip_value=(0, 200),
cleanup=True,
debug=True
)
Conservative Stripping
Use higher f_value to preserve brain tissue:
skull_CTA(
nii_path="pediatric/child_scan.nii.gz",
output_path="conservative_strip/",
f_value=0.3, # Less aggressive
clip_value=(0, 200),
cleanup=True,
debug=True
)
Custom Intensity Range
Adjust clipping for high-contrast scans:
skull_CTA(
nii_path="high_contrast.nii.gz",
output_path="processed/",
f_value=0.1,
clip_value=(0, 300), # Higher upper bound
cleanup=True,
debug=True
)
Quality Control Inspection
Keep intermediates for verification:
skull_CTA(
nii_path="qa/test_scan.nii.gz",
output_path="qa/processing_steps/",
f_value=0.1,
clip_value=(0, 200),
cleanup=False, # Keep all intermediate files
debug=True
)
# Now you can inspect:
# - test_scan_th.nii.gz (after initial threshold)
# - test_scan_th_sm.nii.gz (after smoothing)
# - test_scan_th_sm_th.nii.gz (after second threshold)
# - test_scan_skulled.nii.gz (BET output)
# - test_scan_skulled_clipped.nii.gz (final result)
Verifying FSL Installation
Check FSL before processing:
import subprocess
def check_fsl():
"""Verify FSL is properly installed."""
try:
# Check fslmaths
subprocess.run(['fslmaths'], capture_output=True, check=False)
print("✓ fslmaths found")
# Check bet
subprocess.run(['bet'], capture_output=True, check=False)
print("✓ bet found")
return True
except FileNotFoundError as e:
print(f"✗ FSL not found: {e}")
print("Please install FSL and configure environment variables")
return False
# Verify before processing
if check_fsl():
from nidataset.preprocessing import skull_CTA
skull_CTA(
nii_path="scan.nii.gz",
output_path="processed/",
f_value=0.1,
clip_value=(0, 200),
cleanup=True,
debug=True
)
Testing Different f_values
Find optimal skull-stripping threshold:
from nidataset.preprocessing import skull_CTA
import nibabel as nib
import numpy as np
test_f_values = [0.05, 0.1, 0.15, 0.2, 0.3]
test_scan = "test_data/sample.nii.gz"
results = []
for f_val in test_f_values:
output_dir = f"f_value_test/f_{f_val}/"
skull_CTA(
nii_path=test_scan,
output_path=output_dir,
f_value=f_val,
clip_value=(0, 200),
cleanup=True,
debug=True
)
# Analyze brain coverage
mask = nib.load(f"{output_dir}/sample_skulled_mask.nii.gz")
mask_data = mask.get_fdata()
coverage = np.sum(mask_data > 0) / np.prod(mask_data.shape) * 100
results.append({
'f_value': f_val,
'brain_coverage': coverage
})
print(f"f_value={f_val}: coverage={coverage:.1f}%")
print("\nCompare outputs visually to select optimal f_value")
Assessing Skull Removal Quality
Evaluate processing results:
import nibabel as nib
import numpy as np
from nidataset.preprocessing import skull_CTA
# Process
skull_CTA(
nii_path="qa/original.nii.gz",
output_path="qa/processed/",
f_value=0.1,
clip_value=(0, 200),
cleanup=False, # Keep intermediates for analysis
debug=True
)
# Load results
original = nib.load("qa/original.nii.gz")
mask = nib.load("qa/processed/original_skulled_mask.nii.gz")
stripped = nib.load("qa/processed/original_skulled_clipped.nii.gz")
orig_data = original.get_fdata()
mask_data = mask.get_fdata()
stripped_data = stripped.get_fdata()
print("\nQuality Assessment:")
# 1. Brain coverage
brain_voxels = np.sum(mask_data > 0)
total_voxels = np.prod(mask_data.shape)
coverage = (brain_voxels / total_voxels) * 100
print(f" Brain coverage: {coverage:.1f}%")
print(f" Expected range: 20-40% for head scans")
if coverage < 15:
print(f" ⚠️ Warning: Low coverage, try higher f_value")
elif coverage > 50:
print(f" ⚠️ Warning: High coverage, try lower f_value")
else:
print(f" ✓ Coverage looks reasonable")
# 2. Intensity ranges
print(f"\n Original intensity: [{orig_data.min():.1f}, {orig_data.max():.1f}]")
print(f" Stripped intensity: [{stripped_data.min():.1f}, {stripped_data.max():.1f}]")
print(f" Target clipping: [0, 200]")
# 3. Check masking
brain_region = stripped_data[mask_data > 0]
print(f"\n Brain region mean: {brain_region.mean():.1f}")
print(f" Brain region std: {brain_region.std():.1f}")
Creating Visualization Overlay
Generate overlay for visual inspection:
import nibabel as nib
import numpy as np
from nidataset.preprocessing import skull_CTA
# Process
skull_CTA(
nii_path="visualization/scan.nii.gz",
output_path="visualization/processed/",
f_value=0.1,
clip_value=(0, 200),
cleanup=False,
debug=True
)
# Load original and mask
original = nib.load("visualization/scan.nii.gz")
mask = nib.load("visualization/processed/scan_skulled_mask.nii.gz")
orig_data = original.get_fdata()
mask_data = mask.get_fdata()
# Create edge overlay
from scipy import ndimage
edges = ndimage.sobel(mask_data)
overlay = orig_data.copy()
overlay[edges > 0] = orig_data.max() # Highlight mask boundary
# Save overlay
overlay_img = nib.Nifti1Image(overlay, original.affine)
nib.save(overlay_img, "visualization/mask_overlay.nii.gz")
print("Mask overlay created for quality control")
Error Handling
Robust processing with error management:
from nidataset.preprocessing import skull_CTA
def process_with_error_handling(input_path, output_path):
"""Process scan with comprehensive error handling."""
try:
skull_CTA(
nii_path=input_path,
output_path=output_path,
f_value=0.1,
clip_value=(0, 200),
cleanup=True,
debug=True
)
print(f"✓ Successfully processed: {input_path}")
return True
except FileNotFoundError as e:
print(f"✗ File not found: {e}")
return False
except RuntimeError as e:
print(f"✗ FSL execution failed: {e}")
print("Check FSL installation and environment variables")
return False
except Exception as e:
print(f"✗ Unexpected error: {e}")
return False
# Use in processing
process_with_error_handling(
"data/scan.nii.gz",
"data/processed/"
)
Integration with Vessel Analysis
Complete preprocessing pipeline:
from nidataset.preprocessing import skull_CTA, mip
# Step 1: Skull stripping
print("Step 1: Removing skull...")
skull_CTA(
nii_path="pipeline/original.nii.gz",
output_path="pipeline/skull_stripped/",
f_value=0.1,
clip_value=(0, 200),
cleanup=True,
debug=True
)
# Step 2: Enhance vessels with MIP
print("\nStep 2: Enhancing vessels...")
mip(
nii_path="pipeline/skull_stripped/original_skulled_clipped.nii.gz",
output_path="pipeline/vessel_enhanced/",
window_size=15,
view="axial",
debug=True
)
print("\nPipeline complete: Ready for vessel segmentation")
Batch Processing Helper
Process multiple scans:
from nidataset.preprocessing import skull_CTA
import os
def batch_skull_strip(input_folder, output_base, f_value=0.1):
"""Process all scans in a folder."""
success_count = 0
fail_count = 0
for filename in os.listdir(input_folder):
if filename.endswith('.nii.gz'):
input_path = os.path.join(input_folder, filename)
scan_name = filename.replace('.nii.gz', '')
output_path = os.path.join(output_base, scan_name)
print(f"\nProcessing: {scan_name}")
try:
skull_CTA(
nii_path=input_path,
output_path=output_path,
f_value=f_value,
clip_value=(0, 200),
cleanup=True,
debug=False
)
success_count += 1
print(f"✓ Success")
except Exception as e:
fail_count += 1
print(f"✗ Failed: {e}")
print(f"\n{'='*50}")
print(f"Batch complete: {success_count} success, {fail_count} failed")
# Use
batch_skull_strip("raw_scans/", "processed_scans/", f_value=0.1)
Typical Workflow
from nidataset.preprocessing import skull_CTA
import subprocess
import nibabel as nib
import numpy as np
# 1. Verify FSL installation
try:
subprocess.run(['bet', '--version'], capture_output=True, check=True)
print("✓ FSL is properly configured")
except:
print("✗ Error: FSL not found")
print("Install FSL from https://fsl.fmrib.ox.ac.uk/")
exit(1)
# 2. Process angiography scan
skull_CTA(
nii_path="data/angiography.nii.gz",
output_path="data/skull_stripped/",
f_value=0.1, # Adjust based on scan
clip_value=(0, 200),
cleanup=True,
debug=True
)
# 3. Verify results
mask = nib.load("data/skull_stripped/angiography_skulled_mask.nii.gz")
stripped = nib.load("data/skull_stripped/angiography_skulled_clipped.nii.gz")
mask_data = mask.get_fdata()
coverage = np.sum(mask_data > 0) / np.prod(mask_data.shape) * 100
print(f"\nQuality Check:")
print(f" Brain coverage: {coverage:.1f}%")
print(f" Output shape: {stripped.shape}")
# 4. Use skull-stripped volume for:
# - Vessel segmentation
# - Aneurysm detection
# - Vascular analysis
# - Registration