register_CTA
Register a single medical imaging volume to a reference template using mutual information-based rigid registration with advanced preprocessing.
register_CTA(
nii_path: str,
mask_path: str,
template_path: str,
template_mask_path: str,
output_path: str,
cleanup: bool = False,
debug: bool = False,
number_histogram_bins: int = 128,
learning_rate: float = 0.0001,
number_iterations: int = 2000,
initialization_strategy: int = sitk.CenteredTransformInitializerFilter.MOMENTS,
sigma_first: float = 2.0,
sigma_second: float = 3.0,
metric_sampling_percentage: float = 0.5,
initial_transform = None
) -> None
Overview
This function aligns a medical imaging volume to a reference template through an optimized registration pipeline that combines preprocessing, masking, and mutual information-based optimization. The process ensures robust alignment even with intensity variations and noise.
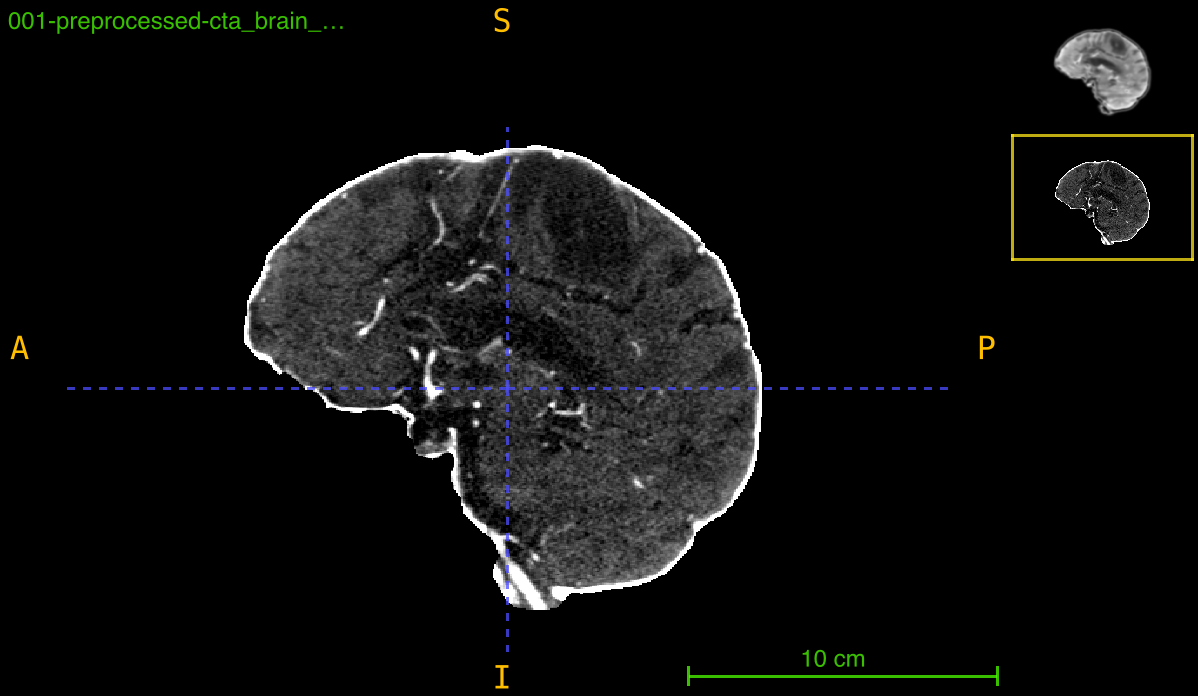
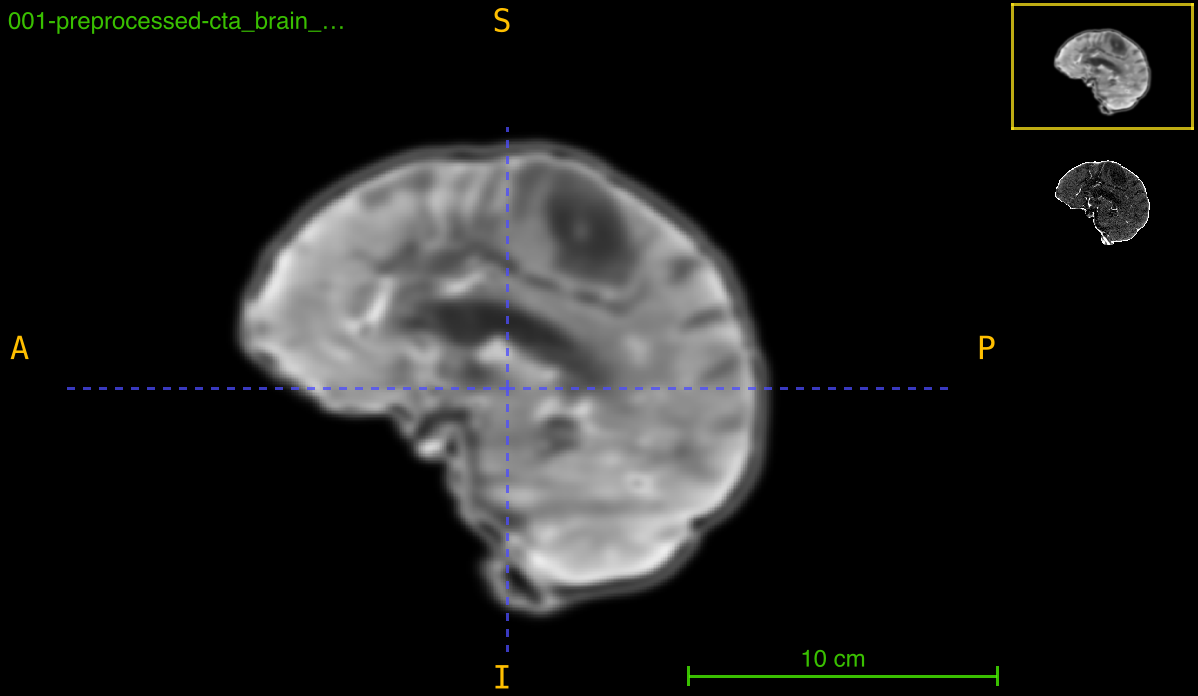
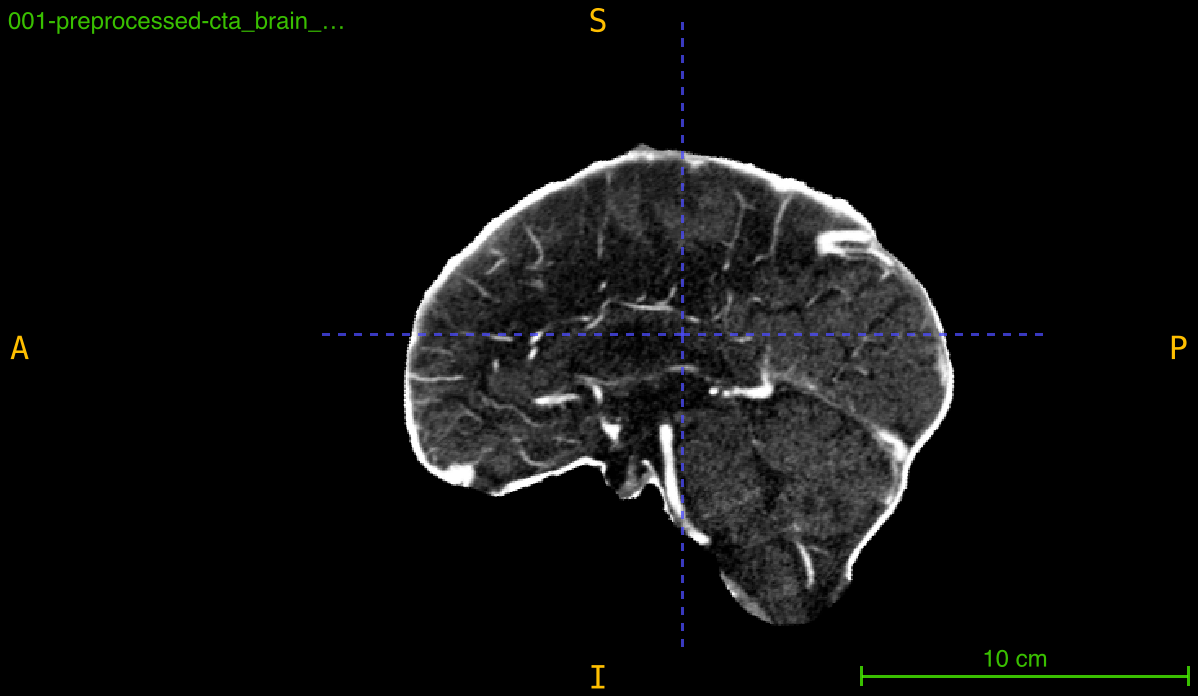
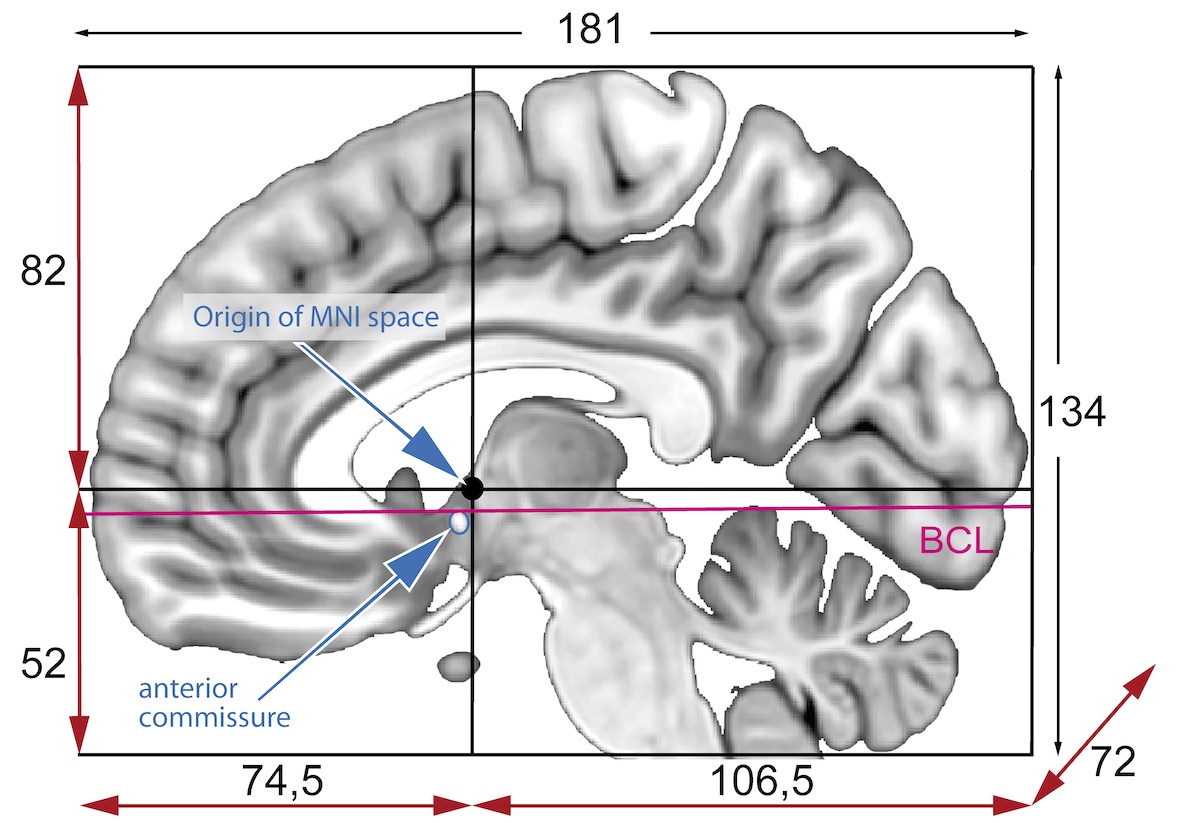
Registration pipeline:
- Preprocessing: Sequential Gaussian filtering with intensity clipping
- Initialization: Moment-based or geometry-based alignment for initial positioning
- Optimization: Gradient descent with mutual information metric
- Transform saving: Parameters stored for reapplication or analysis
This is essential for:
- Standardizing spatial orientation across subjects
- Enabling voxel-wise comparisons
- Creating normalized datasets for analysis
- Applying consistent spatial coordinates
- Building population-based atlases
Parameters
| Name | Type | Default | Description |
|---|---|---|---|
nii_path | str | required | Path to the input volume in .nii.gz format. |
mask_path | str | required | Path to the brain mask for the input volume. |
template_path | str | required | Path to the reference template volume. |
template_mask_path | str | required | Path to the template’s brain mask. |
output_path | str | required | Directory where registered volume and intermediate files will be saved. |
cleanup | bool | False | If True, deletes intermediate Gaussian-filtered file after registration. |
debug | bool | False | If True, prints detailed information about saved file paths. |
number_histogram_bins | int | 128 | Number of histogram bins for Mattes Mutual Information metric. Common values: 10, 50, 64, 128. |
learning_rate | float | 0.0001 | Learning rate for Gradient Descent optimizer. Common values: 0.0001-1.0. |
number_iterations | int | 2000 | Maximum number of optimization iterations. Common values: 500-5000. |
initialization_strategy | int | MOMENTS | Strategy for initializing transformation: MOMENTS (center of mass) or GEOMETRY (center/orientation). |
sigma_first | float | 2.0 | Standard deviation for the first Gaussian smoothing filter. |
sigma_second | float | 3.0 | Standard deviation for the second Gaussian smoothing filter. |
metric_sampling_percentage | float | 0.5 | Percentage of voxels to sample for metric evaluation (0.0-1.0). Default: 0.5 (50%). |
initial_transform | None | None | Initial transformation object. If None, defaults to sitk.Euler3DTransform(). |
Returns
None — The function saves registered volume and transformation to disk.
Output Files
The function generates three files:
| File | Description | Kept After Cleanup |
|---|---|---|
<PREFIX>_registered.nii.gz | Volume aligned to template space | Yes |
<PREFIX>_gaussian_filtered.nii.gz | Preprocessed volume used for registration | No (if cleanup=True) |
<PREFIX>_transformation.tfm | Transformation parameters | Yes |
Example: Input scan_042.nii.gz produces:
scan_042_registered.nii.gzscan_042_gaussian_filtered.nii.gz(temporary)scan_042_transformation.tfm
Preprocessing Pipeline
The volume undergoes multi-stage preprocessing before registration:
Step 1: Negative Value Removal
image[image < 0] = 0
Eliminates artifacts from reconstruction or air regions.
Step 2: Initial Gaussian Smoothing
image = gaussian_filter(image, sigma=sigma_first) # default: 2.0
Reduces noise while preserving structural information.
Step 3: High Intensity Clipping
image[image > 95] = 0
Removes extreme intensities (bone, metal artifacts, or contrast pooling).
Step 4: Secondary Gaussian Smoothing
image = gaussian_filter(image, sigma=sigma_second) # default: 3.0
Further smooths for robust feature matching.
Step 5: Final Intensity Clipping
image = Clamp(image, lowerBound=0, upperBound=100)
Normalizes intensity range to [0, 100] for consistent metric calculation.
Registration Method Details
Initialization
Method: Centered Transform Initializer
- Strategy Options:
MOMENTS(default): Aligns centers of mass between volumes using mask-based moment calculationGEOMETRY: Aligns based on image geometry (center and orientation)
- Provides robust starting point for optimization
Metric
Type: Mattes Mutual Information
- Histogram bins: Configurable (default: 128) for discrete intensity approximation
- Sampling strategy: Random sampling at configurable percentage (default: 50% of voxels)
- Masking: Constrained to brain regions only (both fixed and moving masks)
- Advantage: Robust to intensity variations and scanner differences
Optimization
Algorithm: Gradient Descent
- Learning rate: Configurable (default: 0.0001), estimated once at start
- Iterations: Configurable maximum (default: 2000)
- Scaling: Physical shift-based for balanced optimization
- Convergence: Automatic when improvement plateaus
Transform Type
Model: Euler3D (Rigid transformation) by default, customizable
- 6 degrees of freedom: 3 rotations + 3 translations
- Preserves shape and size
- Suitable for inter-subject brain alignment
- Can be replaced with other transform types via
initial_transformparameter
Interpolation
Method: Linear interpolation
- Balances speed and quality
- Sufficient for most medical imaging applications
Exceptions
| Exception | Condition |
|---|---|
FileNotFoundError | Any required input file does not exist |
ValueError | Input file is not in .nii.gz format |
Usage Notes
- Input Format: Only
.nii.gzfiles are accepted - 3D Volumes Required: All inputs must be 3D NIfTI images
- Mask Requirement: Both volume and template need corresponding masks
- Output Directories: Automatically created if they don’t exist
- Transform Format: SimpleITK
.tfmformat (reusable) - Original Volume: Final registration uses original (not filtered) volume
- Parameter Tuning: Registration quality can be improved by adjusting histogram bins, learning rate, iterations, and sampling percentage
Examples
Basic Usage
Register a single volume to a template:
from nidataset.preprocessing import register_CTA
register_CTA(
nii_path="scans/patient_001.nii.gz",
mask_path="masks/patient_001_mask.nii.gz",
template_path="atlas/standard_template.nii.gz",
template_mask_path="atlas/standard_mask.nii.gz",
output_path="registered/",
cleanup=False,
debug=True
)
# Prints:
# Registered image saved at: 'registered/patient_001_registered.nii.gz'
# Transformation file saved at: 'registered/patient_001_transformation.tfm'
Custom Registration Parameters
Fine-tune registration for better quality:
import SimpleITK as sitk
from nidataset.preprocessing import register_CTA
register_CTA(
nii_path="scans/patient_001.nii.gz",
mask_path="masks/patient_001_mask.nii.gz",
template_path="atlas/standard_template.nii.gz",
template_mask_path="atlas/standard_mask.nii.gz",
output_path="registered/",
number_histogram_bins=64,
learning_rate=0.01,
number_iterations=1000,
initialization_strategy=sitk.CenteredTransformInitializerFilter.GEOMETRY,
sigma_first=1.5,
sigma_second=2.5,
metric_sampling_percentage=0.7,
debug=True
)
Using Custom Initial Transform
Start with an affine transform instead of rigid:
import SimpleITK as sitk
from nidataset.preprocessing import register_CTA
# Create a custom initial transform
affine_transform = sitk.AffineTransform(3)
register_CTA(
nii_path="scans/patient_001.nii.gz",
mask_path="masks/patient_001_mask.nii.gz",
template_path="atlas/standard_template.nii.gz",
template_mask_path="atlas/standard_mask.nii.gz",
output_path="registered/",
initial_transform=affine_transform,
debug=True
)
With Cleanup
Remove intermediate files to save disk space:
register_CTA(
nii_path="data/scan.nii.gz",
mask_path="data/scan_mask.nii.gz",
template_path="template.nii.gz",
template_mask_path="template_mask.nii.gz",
output_path="output/",
cleanup=True, # Removes Gaussian-filtered intermediate
debug=True
)
# Only registered volume and transformation are kept
Quality Control Verification
Register and verify alignment:
import nibabel as nib
import numpy as np
from nidataset.preprocessing import register_CTA
# Perform registration
register_CTA(
nii_path="qa/test_scan.nii.gz",
mask_path="qa/test_mask.nii.gz",
template_path="qa/template.nii.gz",
template_mask_path="qa/template_mask.nii.gz",
output_path="qa/output/",
debug=True
)
# Load results
template = nib.load("qa/template.nii.gz")
registered = nib.load("qa/output/test_scan_registered.nii.gz")
template_data = template.get_fdata()
registered_data = registered.get_fdata()
# Verify alignment
print(f"\nQuality Check:")
print(f" Template shape: {template_data.shape}")
print(f" Registered shape: {registered_data.shape}")
print(f" Shapes match: {template_data.shape == registered_data.shape}")
# Calculate correlation within brain region
mask = nib.load("qa/template_mask.nii.gz").get_fdata()
mask_indices = mask > 0
template_roi = template_data[mask_indices]
registered_roi = registered_data[mask_indices]
correlation = np.corrcoef(template_roi, registered_roi)[0, 1]
print(f" Brain region correlation: {correlation:.3f}")
print(f" Good alignment: {correlation > 0.7}")
Inspecting Preprocessing
Examine intermediate preprocessing steps:
from nidataset.preprocessing import register_CTA
import nibabel as nib
import matplotlib.pyplot as plt
# Register without cleanup to inspect intermediate
register_CTA(
nii_path="inspection/scan.nii.gz",
mask_path="inspection/scan_mask.nii.gz",
template_path="template.nii.gz",
template_mask_path="template_mask.nii.gz",
output_path="inspection/output/",
cleanup=False, # Keep intermediate
debug=True
)
# Load original and filtered
original = nib.load("inspection/scan.nii.gz").get_fdata()
filtered = nib.load("inspection/output/scan_gaussian_filtered.nii.gz").get_fdata()
# Compare middle slices
mid_slice = original.shape[2] // 2
fig, axes = plt.subplots(1, 2, figsize=(12, 5))
axes[0].imshow(original[:, :, mid_slice], cmap='gray')
axes[0].set_title('Original')
axes[1].imshow(filtered[:, :, mid_slice], cmap='gray')
axes[1].set_title('After Gaussian Preprocessing')
plt.savefig('preprocessing_comparison.png')
print("Comparison saved: preprocessing_comparison.png")
Applying Transformation to Other Data
Reuse transformation for related scans:
import SimpleITK as sitk
from nidataset.preprocessing import register_CTA
# Step 1: Register structural scan
register_CTA(
nii_path="structural.nii.gz",
mask_path="structural_mask.nii.gz",
template_path="template.nii.gz",
template_mask_path="template_mask.nii.gz",
output_path="registered/",
debug=True
)
# Step 2: Apply same transformation to functional scan
transform = sitk.ReadTransform("registered/structural_transformation.tfm")
template = sitk.ReadImage("template.nii.gz")
functional = sitk.ReadImage("functional.nii.gz")
# Apply transformation
registered_functional = sitk.Resample(
functional,
template,
transform,
sitk.sitkLinear,
0.0
)
# Save
sitk.WriteImage(registered_functional, "registered/functional_registered.nii.gz")
print("Functional scan registered using structural transformation")
Analyzing Transformation Parameters
Extract and interpret registration parameters:
import SimpleITK as sitk
import numpy as np
from nidataset.preprocessing import register_CTA
# Perform registration
register_CTA(
nii_path="analysis/scan.nii.gz",
mask_path="analysis/scan_mask.nii.gz",
template_path="template.nii.gz",
template_mask_path="template_mask.nii.gz",
output_path="analysis/output/",
debug=True
)
# Load transformation
transform = sitk.ReadTransform("analysis/output/scan_transformation.tfm")
params = transform.GetParameters()
print("\nTransformation Analysis:")
print(f" Transform type: {transform.GetName()}")
print(f" Number of parameters: {len(params)}")
# For Euler3D transform (6 parameters: 3 rotations + 3 translations)
if len(params) >= 6:
rotations = params[0:3] # Radians
translations = params[3:6] # mm
print(f"\n Rotations (radians):")
print(f" X-axis: {rotations[0]:.4f} ({np.degrees(rotations[0]):.2f}°)")
print(f" Y-axis: {rotations[1]:.4f} ({np.degrees(rotations[1]):.2f}°)")
print(f" Z-axis: {rotations[2]:.4f} ({np.degrees(rotations[2]):.2f}°)")
print(f"\n Translations (mm):")
print(f" X: {translations[0]:.2f}")
print(f" Y: {translations[1]:.2f}")
print(f" Z: {translations[2]:.2f}")
total_translation = np.sqrt(sum(t**2 for t in translations))
print(f"\n Total translation: {total_translation:.2f} mm")
Batch Processing with Error Handling
Process multiple volumes robustly:
from nidataset.preprocessing import register_CTA
import os
scans = ["scan_001.nii.gz", "scan_002.nii.gz", "scan_003.nii.gz"]
template = "atlas/template.nii.gz"
template_mask = "atlas/template_mask.nii.gz"
failed_scans = []
for scan_file in scans:
scan_id = scan_file.replace('.nii.gz', '')
try:
register_CTA(
nii_path=f"scans/{scan_file}",
mask_path=f"masks/{scan_file}",
template_path=template,
template_mask_path=template_mask,
output_path="registered/",
cleanup=True,
debug=True
)
print(f"✓ Successfully registered: {scan_id}")
except Exception as e:
print(f"✗ Failed: {scan_id} - {str(e)}")
failed_scans.append(scan_id)
if failed_scans:
print(f"\nFailed scans ({len(failed_scans)}):")
for scan in failed_scans:
print(f" - {scan}")
Comparing Registration Quality
Register with different parameters to assess impact:
import nibabel as nib
import numpy as np
from nidataset.preprocessing import register_CTA
# Standard registration
register_CTA(
nii_path="comparison/scan.nii.gz",
mask_path="comparison/scan_mask.nii.gz",
template_path="template.nii.gz",
template_mask_path="template_mask.nii.gz",
output_path="comparison/standard/",
cleanup=False,
debug=True
)
# High-quality registration
register_CTA(
nii_path="comparison/scan.nii.gz",
mask_path="comparison/scan_mask.nii.gz",
template_path="template.nii.gz",
template_mask_path="template_mask.nii.gz",
output_path="comparison/high_quality/",
number_histogram_bins=256,
learning_rate=0.001,
number_iterations=5000,
metric_sampling_percentage=0.8,
cleanup=False,
debug=True
)
# Load and compare results
template_data = nib.load("template.nii.gz").get_fdata()
template_mask = nib.load("template_mask.nii.gz").get_fdata()
registered_standard = nib.load("comparison/standard/scan_registered.nii.gz").get_fdata()
registered_hq = nib.load("comparison/high_quality/scan_registered.nii.gz").get_fdata()
# Calculate metrics within brain
mask_indices = template_mask > 0
template_roi = template_data[mask_indices]
standard_roi = registered_standard[mask_indices]
hq_roi = registered_hq[mask_indices]
# Correlations
corr_standard = np.corrcoef(template_roi, standard_roi)[0, 1]
corr_hq = np.corrcoef(template_roi, hq_roi)[0, 1]
print(f"\nRegistration Quality Comparison:")
print(f" Standard correlation: {corr_standard:.3f}")
print(f" High-quality correlation: {corr_hq:.3f}")
print(f" Improvement: {corr_hq - corr_standard:.3f}")
Integration with Pipeline
Use registration in a complete preprocessing workflow:
from nidataset.preprocessing import register_CTA
from nidataset.volume import generate_brain_mask, crop_and_pad
import os
def preprocess_scan(scan_path, template_path, template_mask_path, output_folder):
"""Complete preprocessing pipeline with registration."""
scan_id = os.path.basename(scan_path).replace('.nii.gz', '')
# Step 1: Generate brain mask
print(f"Step 1: Generating brain mask for {scan_id}...")
generate_brain_mask(
nii_path=scan_path,
output_path=f"{output_folder}/masks/",
threshold=(50, 300),
closing_radius=3
)
# Step 2: Register to template
print(f"Step 2: Registering {scan_id}...")
mask_path = f"{output_folder}/masks/{scan_id}_brain_mask.nii.gz"
register_CTA(
nii_path=scan_path,
mask_path=mask_path,
template_path=template_path,
template_mask_path=template_mask_path,
output_path=f"{output_folder}/registered/",
cleanup=True,
debug=True
)
# Step 3: Crop and pad registered volume
print(f"Step 3: Cropping and padding {scan_id}...")
registered_path = f"{output_folder}/registered/{scan_id}_registered.nii.gz"
crop_and_pad(
nii_path=registered_path,
output_path=f"{output_folder}/final/",
target_shape=(128, 128, 128)
)
print(f"Preprocessing complete for {scan_id}")
# Use in pipeline
preprocess_scan(
scan_path="raw/patient_001.nii.gz",
template_path="atlas/template.nii.gz",
template_mask_path="atlas/template_mask.nii.gz",
output_folder="preprocessed/"
)
Typical Workflow
from nidataset.preprocessing import register_CTA
import nibabel as nib
# 1. Prepare inputs
scan_file = "data/patient_scan.nii.gz"
scan_mask = "data/patient_mask.nii.gz"
template = "atlas/standard_template.nii.gz"
template_mask = "atlas/standard_mask.nii.gz"
# 2. Perform registration
register_CTA(
nii_path=scan_file,
mask_path=scan_mask,
template_path=template,
template_mask_path=template_mask,
output_path="registered/",
cleanup=True, # Save disk space
debug=True
)
# 3. Verify result
registered = nib.load("registered/patient_scan_registered.nii.gz")
template_img = nib.load(template)
print(f"Template shape: {template_img.shape}")
print(f"Registered shape: {registered.shape}")
# 4. Use registered volume for:
# - Voxel-wise analysis
# - Population studies
# - Group comparisons
# - Atlas-based segmentation
Parameter Tuning Guide
| Parameter | Effect | Recommendations |
|---|---|---|
number_histogram_bins | Higher values = finer intensity discretization | 64-128 for most cases; 256 for high-contrast images |
learning_rate | Higher values = faster but less stable convergence | 0.0001-0.001 for standard; 0.01+ for fast initial alignment |
number_iterations | More iterations = potential for better alignment | 1000-2000 standard; 3000-5000 for difficult cases |
metric_sampling_percentage | Higher sampling = more accurate but slower | 0.3-0.5 for speed; 0.7-1.0 for accuracy |
sigma_first / sigma_second | Controls smoothing strength | Lower for sharp features; higher for noisy images |
initialization_strategy | MOMENTS vs GEOMETRY | MOMENTS for asymmetric anatomy; GEOMETRY for symmetric |