register_annotation
Apply a saved transformation to an annotation and optionally recalculate bounding box around the transformed region.
register_annotation(
annotation_path: str,
transform_path: str,
registered_path: str,
output_path: str,
recalculate_bbox: bool = True,
debug: bool = False
) -> None
Overview
This function transforms an annotation (typically a bounding box mask) using a previously computed registration transformation. It can either preserve the deformed annotation or create a new tight bounding box around the transformed region, which is particularly useful for maintaining axis-aligned boxes after rotation or spatial transformations.
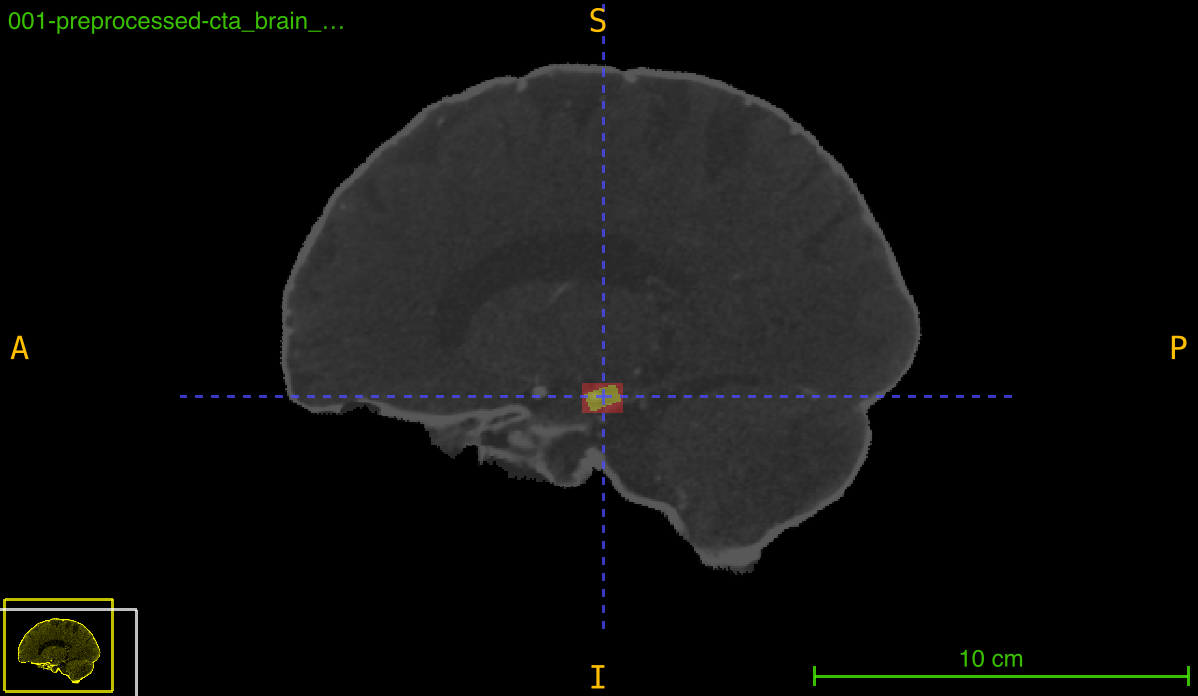
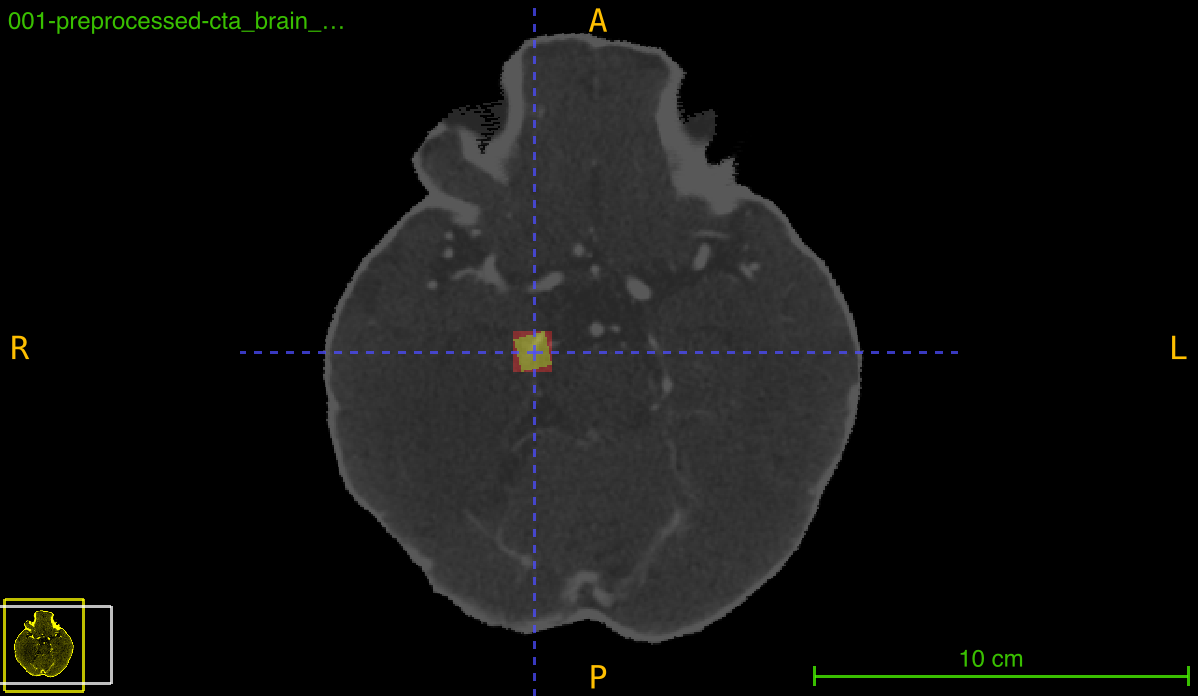
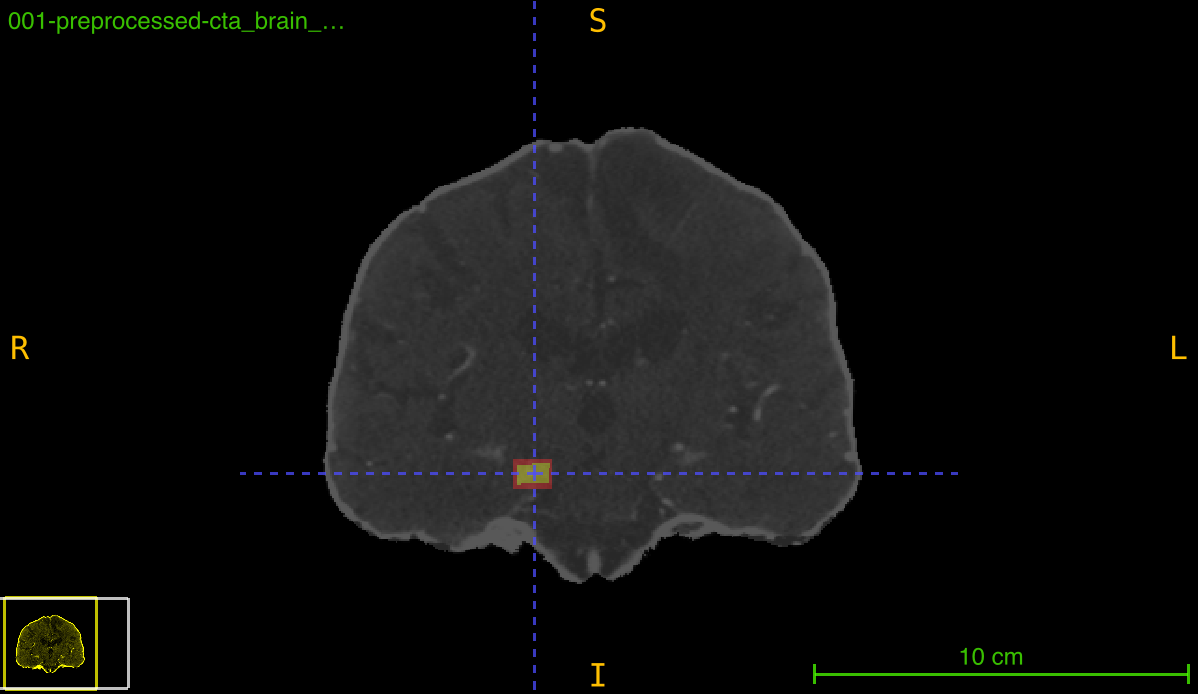
Transformation pipeline:
- Load: Reads annotation, transformation, and reference image
- Transform: Applies transformation using nearest neighbor interpolation
- Recalculate (optional): Computes new axis-aligned bounding box around transformed region
- Output: Saves transformed annotation or recalculated bounding box
This is essential for:
- Maintaining axis-aligned bounding boxes after registration
- Propagating region-of-interest annotations to normalized space
- Ensuring annotations remain tight and efficient after rotation
- Transforming object detection labels for registered volumes
- Creating standardized annotation datasets
Parameters
| Name | Type | Default | Description |
|---|---|---|---|
annotation_path | str | required | Path to the input annotation file ending with _bbox.nii.gz. |
transform_path | str | required | Path to the transformation file (.tfm) from a previous registration. |
registered_path | str | required | Path to the registered image that defines the target space and grid. |
output_path | str | required | Path where the transformed annotation will be saved (including filename). |
recalculate_bbox | bool | True | If True, creates new axis-aligned bounding box. If False, preserves deformed shape. |
debug | bool | False | If True, prints detailed information about the transformation and bounding box dimensions. |
Returns
None — The function saves the transformed annotation to disk.
Output Files
The function generates a single file:
| File | Description | When recalculate_bbox=True | When recalculate_bbox=False |
|---|---|---|---|
| User-specified output path | Transformed annotation | New axis-aligned bounding box | Deformed annotation shape |
Bounding Box Recalculation
With Recalculation (recalculate_bbox=True)
Purpose: Creates a new tight, axis-aligned bounding box around the transformed region.
Process:
- Apply transformation to annotation
- Find all non-zero voxels in transformed space
- Calculate minimum bounding box containing all voxels
- Create new box:
[x_min:x_max, y_min:y_max, z_min:z_max]
Advantages:
- Maintains axis alignment (important for many detection algorithms)
- Ensures tight bounding around transformed region
- Removes any deformation artifacts
- Produces clean rectangular boxes
Use when:
- Working with object detection frameworks
- Need axis-aligned boxes for downstream processing
- Rotation was applied during registration
- Box efficiency matters more than exact shape
Visual Example:
Original Box: After Rotation: After Recalculation:
┌───────────┐ ╱───────────╲ ┌────────────┐
│ │ ╱ ╲ │ ╱────────╲ │
│ Region │ → ╱ Region ╲ → │╱ Region ╲│
│ │ ╲ ╱ │╲──────────╱│
└───────────┘ ╲─────────────╱ └────────────┘
(Deformed) (Tight Box)
Without Recalculation (recalculate_bbox=False)
Purpose: Preserves the exact deformed shape after transformation.
Process:
- Apply transformation to annotation
- Save the transformed shape as-is
Advantages:
- Preserves exact spatial deformation
- More accurate for shape-critical applications
- Maintains relative positions within annotation
Use when:
- Exact deformed shape is important
- Working with precise spatial relationships
- Annotation is not a simple bounding box
- Deformation information needs preservation
Important Notes
Annotation Naming Convention
- All annotation files must end with
_bbox.nii.gz - This is enforced by the function’s validation
- Examples of valid names:
patient001_lesion_bbox.nii.gzcase001_tumor_bbox.nii.gzvessel_roi_bbox.nii.gz
- Examples of invalid names:
patient001_lesion.nii.gz(missing_bbox)lesion.nii.gz(missing_bbox)bbox_lesion.nii.gz(_bboxnot at end before.nii.gz)
File Requirements
- Input annotation must be a 3D NIfTI image (
.nii.gz) - Transformation file must exist from a prior registration
- Reference image defines the target coordinate space
- All three input files must exist before function call
Exceptions
| Exception | Condition |
|---|---|
FileNotFoundError | Any required input file (annotation, transform, or reference) does not exist |
ValueError | Input annotation filename does not end with _bbox.nii.gz |
Usage Notes
- Input Format: Only
.nii.gzfiles ending with_bboxare accepted - 3D Annotations Required: Input must be 3D NIfTI image
- Transform Dependency: Transformation must be from a completed registration
- Empty Annotations: If transformation results in empty annotation (no non-zero voxels), saves empty mask with warning
- Interpolation: Always uses nearest neighbor to preserve binary annotation values
- Output Directories: Automatically created if they don’t exist
- Coordinate System: Output coordinates are in the reference image space
Examples
Basic Usage - Recalculate Bounding Box
Transform annotation and create new axis-aligned box:
from nidataset.preprocessing import register_CTA, register_annotation
# Step 1: Register the CTA scan
register_CTA(
nii_path="scan.nii.gz",
mask_path="scan_mask.nii.gz",
template_path="template.nii.gz",
template_mask_path="template_mask.nii.gz",
output_path="registered/",
debug=True
)
# Step 2: Transform bounding box annotation with recalculation
register_annotation(
annotation_path="scan_lesion_bbox.nii.gz",
transform_path="registered/scan_transformation.tfm",
registered_path="registered/scan_registered.nii.gz",
output_path="registered/scan_lesion_bbox_registered.nii.gz",
recalculate_bbox=True,
debug=True
)
# Prints:
# Bounding box recalculated:
# Original bbox size: 15847 voxels
# New bbox: [45:98, 67:112, 34:78]
# Registered annotation saved at: 'registered/scan_lesion_bbox_registered.nii.gz'
Preserve Deformed Shape
Keep exact transformed annotation shape:
from nidataset.preprocessing import register_annotation
# Transform without recalculation - preserves deformation
register_annotation(
annotation_path="scan_region_bbox.nii.gz",
transform_path="registered/scan_transformation.tfm",
registered_path="registered/scan_registered.nii.gz",
output_path="registered/scan_region_registered.nii.gz",
recalculate_bbox=False, # Keep deformed shape
debug=True
)
# Prints:
# Registered annotation saved at: 'registered/scan_region_registered.nii.gz'
Transform Multiple Annotations
Process several annotations from the same scan:
from nidataset.preprocessing import register_CTA, register_annotation
import os
# Register scan once
register_CTA(
nii_path="patient001.nii.gz",
mask_path="patient001_mask.nii.gz",
template_path="template.nii.gz",
template_mask_path="template_mask.nii.gz",
output_path="registered/patient001/"
)
# Transform multiple bounding box annotations
annotations = [
"patient001_lesion1_bbox.nii.gz",
"patient001_lesion2_bbox.nii.gz",
"patient001_vessel_bbox.nii.gz",
"patient001_hemorrhage_bbox.nii.gz"
]
for annotation_file in annotations:
output_name = annotation_file.replace("_bbox.nii.gz", "_bbox_registered.nii.gz")
register_annotation(
annotation_path=f"annotations/{annotation_file}",
transform_path="registered/patient001/patient001_transformation.tfm",
registered_path="registered/patient001/patient001_registered.nii.gz",
output_path=f"registered/patient001/{output_name}",
recalculate_bbox=True,
debug=True
)
print(f"Registered: {annotation_file}")
Compare Recalculation vs Preservation
Evaluate the difference between both modes:
from nidataset.preprocessing import register_annotation
import nibabel as nib
import numpy as np
# Transform with recalculation
register_annotation(
annotation_path="lesion_bbox.nii.gz",
transform_path="transforms/transformation.tfm",
registered_path="registered/scan_registered.nii.gz",
output_path="comparison/bbox_recalculated.nii.gz",
recalculate_bbox=True
)
# Transform without recalculation
register_annotation(
annotation_path="lesion_bbox.nii.gz",
transform_path="transforms/transformation.tfm",
registered_path="registered/scan_registered.nii.gz",
output_path="comparison/bbox_deformed.nii.gz",
recalculate_bbox=False
)
# Load and compare
recalc = nib.load("comparison/bbox_recalculated.nii.gz").get_fdata()
deformed = nib.load("comparison/bbox_deformed.nii.gz").get_fdata()
print("\nComparison:")
print(f" Recalculated volume: {np.sum(recalc > 0)} voxels")
print(f" Deformed volume: {np.sum(deformed > 0)} voxels")
print(f" Overlap: {np.sum((recalc > 0) & (deformed > 0))} voxels")
print(f" Volume difference: {abs(np.sum(recalc > 0) - np.sum(deformed > 0))} voxels")
# Recalculated is typically larger (encompasses all deformed voxels)
# but maintains axis alignment
Batch Process Annotations
Transform all annotations in a dataset:
from nidataset.preprocessing import register_annotation
import os
from tqdm import tqdm
def batch_register_annotations(annotation_folder, transform_folder,
registered_folder, output_path,
recalculate_bbox=True):
"""Register all annotations in a folder."""
annotation_files = [f for f in os.listdir(annotation_folder)
if f.endswith("_bbox.nii.gz")]
os.makedirs(output_path, exist_ok=True)
success_count = 0
failed_cases = []
for annotation_file in tqdm(annotation_files, desc="Registering annotations"):
# Extract prefix by removing _bbox.nii.gz
prefix = annotation_file.replace("_bbox.nii.gz", "")
# Find corresponding transform and reference
transform_file = os.path.join(transform_folder, f"{prefix}_transformation.tfm")
reference_file = os.path.join(registered_folder, f"{prefix}_registered.nii.gz")
if not os.path.exists(transform_file) or not os.path.exists(reference_file):
failed_cases.append((prefix, "Missing files"))
continue
try:
output_file = os.path.join(output_path,
f"{prefix}_bbox_registered.nii.gz")
register_annotation(
annotation_path=os.path.join(annotation_folder, annotation_file),
transform_path=transform_file,
registered_path=reference_file,
output_path=output_file,
recalculate_bbox=recalculate_bbox,
debug=False
)
success_count += 1
except Exception as e:
failed_cases.append((prefix, str(e)))
print(f"\nBatch Registration Summary:")
print(f" Total: {len(annotation_files)}")
print(f" Success: {success_count}")
print(f" Failed: {len(failed_cases)}")
if failed_cases:
print("\nFailed cases:")
for case, reason in failed_cases:
print(f" - {case}: {reason}")
return success_count, failed_cases
# Use batch processing
batch_register_annotations(
annotation_folder="data/bboxes/",
transform_folder="data/registered/transforms/",
registered_folder="data/registered/registered/",
output_path="data/bboxes_registered/",
recalculate_bbox=True
)
Extract Bounding Box Coordinates
Get transformed bounding box coordinates for analysis:
from nidataset.preprocessing import register_annotation
import nibabel as nib
import numpy as np
# Register annotation with recalculation
register_annotation(
annotation_path="lesion_bbox.nii.gz",
transform_path="transforms/transformation.tfm",
registered_path="registered/scan_registered.nii.gz",
output_path="registered/lesion_bbox_registered.nii.gz",
recalculate_bbox=True,
debug=True
)
# Load registered annotation
registered_bbox = nib.load("registered/lesion_bbox_registered.nii.gz")
bbox_data = registered_bbox.get_fdata()
# Extract bounding box coordinates
coords = np.argwhere(bbox_data > 0)
if coords.size > 0:
z_min, y_min, x_min = coords.min(axis=0)
z_max, y_max, x_max = coords.max(axis=0)
# Calculate dimensions
width = x_max - x_min + 1
height = y_max - y_min + 1
depth = z_max - z_min + 1
volume = width * height * depth
print("\nBounding Box Information:")
print(f" Coordinates: [{x_min}:{x_max}, {y_min}:{y_max}, {z_min}:{z_max}]")
print(f" Dimensions: {width} × {height} × {depth}")
print(f" Volume: {volume} voxels")
print(f" Center: ({(x_min+x_max)/2:.1f}, {(y_min+y_max)/2:.1f}, {(z_min+z_max)/2:.1f})")
else:
print("Warning: Bounding box is empty")
Create Object Detection Dataset
Prepare annotations for machine learning:
from nidataset.preprocessing import register_CTA, register_annotation
import nibabel as nib
import numpy as np
import json
import os
def create_detection_dataset(scan_folder, bbox_folder, output_folder,
template_path, template_mask_path):
"""
Create normalized dataset with registered scans and bounding boxes.
Exports metadata in COCO-like format for object detection.
"""
os.makedirs(f"{output_folder}/scans", exist_ok=True)
os.makedirs(f"{output_folder}/annotations", exist_ok=True)
scan_files = [f for f in os.listdir(scan_folder) if f.endswith(".nii.gz")]
dataset_metadata = {
"images": [],
"annotations": []
}
for scan_file in scan_files:
case_id = scan_file.replace(".nii.gz", "")
print(f"\nProcessing {case_id}...")
# Register scan
register_CTA(
nii_path=os.path.join(scan_folder, scan_file),
mask_path=os.path.join(scan_folder.replace("scans", "masks"),
f"{case_id}_mask.nii.gz"),
template_path=template_path,
template_mask_path=template_mask_path,
output_path=f"{output_folder}/scans/{case_id}/",
cleanup=True
)
# Register bounding box
bbox_file = os.path.join(bbox_folder, f"{case_id}_bbox.nii.gz")
if not os.path.exists(bbox_file):
print(f" Warning: No bounding box found for {case_id}")
continue
register_annotation(
annotation_path=bbox_file,
transform_path=f"{output_folder}/scans/{case_id}/{case_id}_transformation.tfm",
registered_path=f"{output_folder}/scans/{case_id}/{case_id}_registered.nii.gz",
output_path=f"{output_folder}/annotations/{case_id}_bbox_registered.nii.gz",
recalculate_bbox=True,
debug=False
)
# Extract bbox coordinates
bbox_data = nib.load(
f"{output_folder}/annotations/{case_id}_bbox_registered.nii.gz"
).get_fdata()
coords = np.argwhere(bbox_data > 0)
if coords.size > 0:
z_min, y_min, x_min = coords.min(axis=0)
z_max, y_max, x_max = coords.max(axis=0)
# Add to metadata
dataset_metadata["images"].append({
"id": case_id,
"file_name": f"scans/{case_id}/{case_id}_registered.nii.gz"
})
dataset_metadata["annotations"].append({
"image_id": case_id,
"bbox": [int(x_min), int(y_min), int(z_min),
int(x_max), int(y_max), int(z_max)],
"area": int((x_max - x_min) * (y_max - y_min) * (z_max - z_min))
})
print(f" Registered with bbox: [{x_min}:{x_max}, {y_min}:{y_max}, {z_min}:{z_max}]")
# Save metadata
with open(f"{output_folder}/dataset_metadata.json", "w") as f:
json.dump(dataset_metadata, f, indent=2)
print(f"\nDataset created: {len(dataset_metadata['images'])} scans with annotations")
print(f"Metadata saved: {output_folder}/dataset_metadata.json")
# Create detection dataset
create_detection_dataset(
scan_folder="raw_data/scans/",
bbox_folder="raw_data/bboxes/",
output_folder="detection_dataset/",
template_path="atlas/template.nii.gz",
template_mask_path="atlas/template_mask.nii.gz"
)
Visualize Transformed Annotations
Create visual comparison of annotations:
from nidataset.preprocessing import register_annotation
import nibabel as nib
import matplotlib.pyplot as plt
import numpy as np
# Register with recalculation
register_annotation(
annotation_path="lesion_bbox.nii.gz",
transform_path="transforms/transformation.tfm",
registered_path="registered/scan_registered.nii.gz",
output_path="registered/bbox_recalc.nii.gz",
recalculate_bbox=True
)
# Register without recalculation
register_annotation(
annotation_path="lesion_bbox.nii.gz",
transform_path="transforms/transformation.tfm",
registered_path="registered/scan_registered.nii.gz",
output_path="registered/bbox_deformed.nii.gz",
recalculate_bbox=False
)
# Load for visualization
scan = nib.load("registered/scan_registered.nii.gz").get_fdata()
bbox_recalc = nib.load("registered/bbox_recalc.nii.gz").get_fdata()
bbox_deformed = nib.load("registered/bbox_deformed.nii.gz").get_fdata()
# Select middle slice
mid_slice = scan.shape[2] // 2
# Create visualization
fig, axes = plt.subplots(1, 3, figsize=(18, 6))
# Original scan
axes[0].imshow(scan[:, :, mid_slice], cmap='gray')
axes[0].set_title('Registered Scan', fontsize=14)
axes[0].axis('off')
# With recalculated bbox
axes[1].imshow(scan[:, :, mid_slice], cmap='gray')
axes[1].imshow(bbox_recalc[:, :, mid_slice], cmap='Reds', alpha=0.4)
axes[1].set_title('Recalculated BBox\n(Axis-aligned)', fontsize=14)
axes[1].axis('off')
# With deformed bbox
axes[2].imshow(scan[:, :, mid_slice], cmap='gray')
axes[2].imshow(bbox_deformed[:, :, mid_slice], cmap='Blues', alpha=0.4)
axes[2].set_title('Deformed BBox\n(Preserves shape)', fontsize=14)
axes[2].axis('off')
plt.tight_layout()
plt.savefig('annotation_comparison.png', dpi=150, bbox_inches='tight')
print("Visualization saved: annotation_comparison.png")
Handle Empty Annotations
Properly handle cases where transformation results in empty annotation:
from nidataset.preprocessing import register_annotation
import nibabel as nib
import numpy as np
def safe_register_annotation(annotation_path, transform_path,
reference_path, output_path,
recalculate_bbox=True):
"""Register annotation with empty annotation handling."""
# Register annotation
register_annotation(
annotation_path=annotation_path,
transform_path=transform_path,
registered_path=reference_path,
output_path=output_path,
recalculate_bbox=recalculate_bbox,
debug=True
)
# Check if result is empty
result = nib.load(output_path).get_fdata()
if np.sum(result > 0) == 0:
print(f"Warning: Registered annotation is empty")
print(f" This may indicate the annotation was transformed outside the")
print(f" reference image boundaries or registration misalignment.")
return False
else:
print(f"Annotation registered successfully")
print(f" Contains {np.sum(result > 0)} voxels")
return True
# Use safe registration
success = safe_register_annotation(
annotation_path="small_lesion_bbox.nii.gz",
transform_path="transforms/transformation.tfm",
reference_path="registered/scan_registered.nii.gz",
output_path="registered/lesion_bbox_registered.nii.gz"
)
if not success:
print("Consider checking:")
print(" 1. Original annotation location")
print(" 2. Registration quality")
print(" 3. Template coverage")
Integration with Detection Pipeline
Complete workflow for object detection:
from nidataset.preprocessing import register_CTA_dataset, register_annotation
import os
import pandas as pd
def prepare_detection_dataset(base_dir, template_path, template_mask_path):
"""
Prepare a complete object detection dataset.
Expected structure:
base_dir/
├── scans/
├── masks/
└── bounding_boxes/ (files ending with _bbox.nii.gz)
"""
print("=" * 60)
print("STEP 1: Register CTA scans")
print("=" * 60)
# Register all scans
register_CTA_dataset(
nii_folder=f"{base_dir}/scans/",
mask_folder=f"{base_dir}/masks/",
template_path=template_path,
template_mask_path=template_mask_path,
output_path=f"{base_dir}/registered/",
saving_mode="case",
cleanup=True,
debug=True
)
print("\n" + "=" * 60)
print("STEP 2: Transform bounding box annotations")
print("=" * 60)
# Get list of cases
cases = [d for d in os.listdir(f"{base_dir}/registered/")
if os.path.isdir(os.path.join(f"{base_dir}/registered/", d))]
bbox_records = []
for case in cases:
bbox_file = f"{base_dir}/bounding_boxes/{case}_bbox.nii.gz"
if not os.path.exists(bbox_file):
print(f"No bounding box found for {case}")
continue
try:
output_path = f"{base_dir}/registered/{case}/{case}_bbox_registered.nii.gz"
register_annotation(
annotation_path=bbox_file,
transform_path=f"{base_dir}/registered/{case}/{case}_transformation.tfm",
registered_path=f"{base_dir}/registered/{case}/{case}_registered.nii.gz",
output_path=output_path,
recalculate_bbox=True,
debug=False
)
# Extract bbox coordinates
import nibabel as nib
import numpy as np
bbox_data = nib.load(output_path).get_fdata()
coords = np.argwhere(bbox_data > 0)
if coords.size > 0:
z_min, y_min, x_min = coords.min(axis=0)
z_max, y_max, x_max = coords.max(axis=0)
bbox_records.append({
'case_id': case,
'x_min': int(x_min),
'y_min': int(y_min),
'z_min': int(z_min),
'x_max': int(x_max),
'y_max': int(y_max),
'z_max': int(z_max),
'width': int(x_max - x_min + 1),
'height': int(y_max - y_min + 1),
'depth': int(z_max - z_min + 1)
})
print(f"{case}: bbox [{x_min}:{x_max}, {y_min}:{y_max}, {z_min}:{z_max}]")
else:
print(f"{case}: Empty annotation after transformation")
except Exception as e:
print(f"{case}: Failed - {str(e)}")
# Save bounding box catalog
if bbox_records:
df = pd.DataFrame(bbox_records)
catalog_path = f"{base_dir}/bbox_catalog.csv"
df.to_csv(catalog_path, index=False)
print("\n" + "=" * 60)
print("SUMMARY")
print("=" * 60)
print(f"Processed {len(bbox_records)} cases")
print(f"Catalog saved: {catalog_path}")
print("\nBounding box statistics:")
print(f" Mean width: {df['width'].mean():.1f} voxels")
print(f" Mean height: {df['height'].mean():.1f} voxels")
print(f" Mean depth: {df['depth'].mean():.1f} voxels")
# Run complete pipeline
prepare_detection_dataset(
base_dir="detection_project",
template_path="atlas/template.nii.gz",
template_mask_path="atlas/template_mask.nii.gz"
)
Typical Workflow
from nidataset.preprocessing import register_CTA, register_annotation
# Step 1: Register scan
register_CTA(
nii_path="scan.nii.gz",
mask_path="scan_mask.nii.gz",
template_path="template.nii.gz",
template_mask_path="template_mask.nii.gz",
output_path="registered/"
)
# Step 2: Transform bounding box annotations
register_annotation(
annotation_path="scan_lesion_bbox.nii.gz",
transform_path="registered/scan_transformation.tfm",
registered_path="registered/scan_registered.nii.gz",
output_path="registered/scan_lesion_bbox_registered.nii.gz",
recalculate_bbox=True # For axis-aligned boxes
)
# Step 3: Use registered annotations for:
# - Object detection training
# - Region-of-interest analysis
# - Automated lesion detection
# - Bounding box-based segmentation