resampling
Resample a 3D medical imaging volume to a target size while preserving the physical field of view through automatic voxel spacing adjustment.
resampling(
nii_path: str,
output_path: str,
desired_volume: tuple,
debug: bool = False
) -> None
Overview
This function changes the dimensions of a 3D volume to a target size while maintaining the original physical extent (field of view) by automatically calculating and applying new voxel spacing. Unlike simple array resizing, resampling ensures that anatomical structures retain their true physical dimensions.
Key process:
- Calculate new spacing:
new_spacing = old_spacing × (old_size / new_size) - Apply B-spline interpolation: Smooth, high-quality resampling
- Preserve field of view: Physical dimensions remain constant
- Maintain metadata: Origin and direction are preserved
This is essential for:
- Standardizing volume dimensions for deep learning models
- Creating uniform datasets from variable-sized scans
- Reducing computational requirements while preserving anatomy
- Preparing data for fixed-size network inputs
- Balancing detail and processing speed
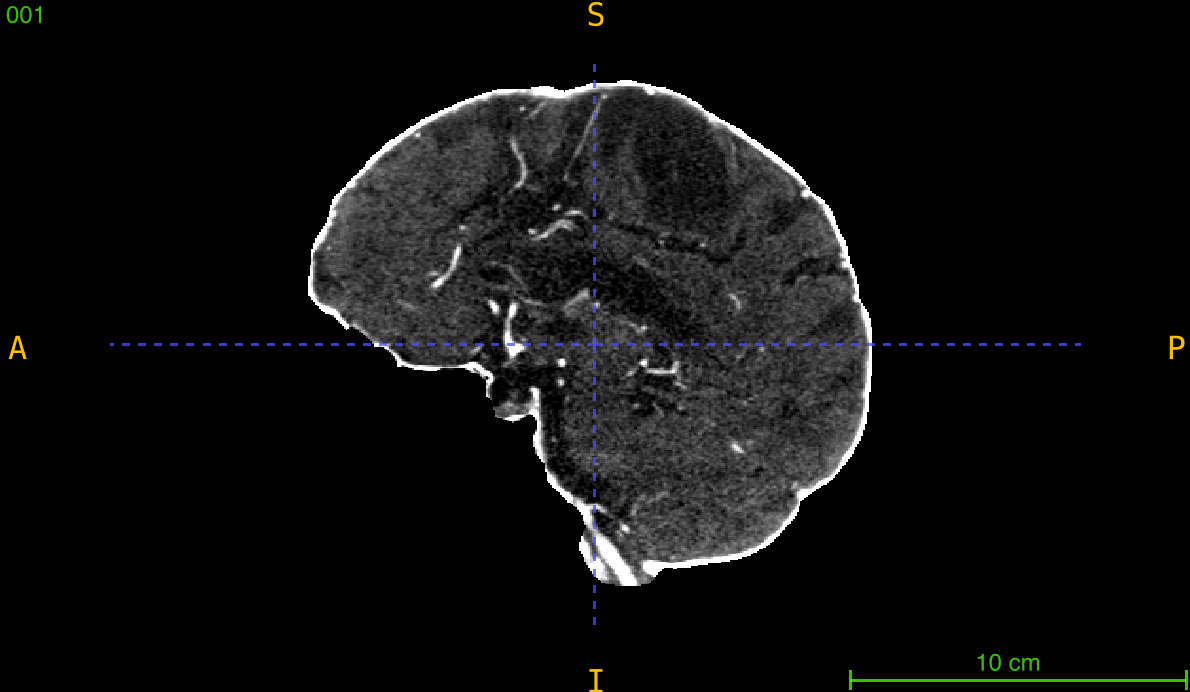
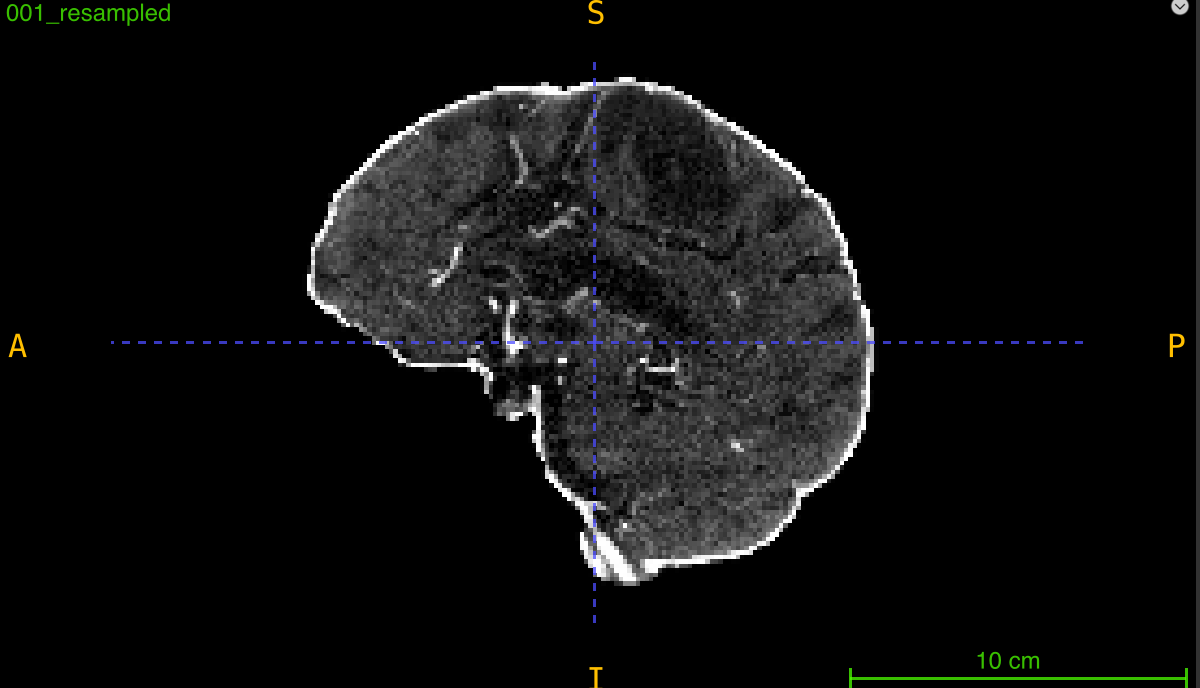
Parameters
| Name | Type | Default | Description |
|---|---|---|---|
nii_path | str | required | Path to the input volume in .nii.gz format. |
output_path | str | required | Directory where the resampled volume will be saved. Created automatically if it doesn’t exist. |
desired_volume | tuple | required | Target volume size as (X, Y, Z). Must contain exactly three integers. |
debug | bool | False | If True, prints the output file path. |
Returns
None – The function saves the resampled volume to disk.
Output File
The resampled volume is saved as:
<PREFIX>_resampled.nii.gz
Example: Input scan_042.nii.gz → Output scan_042_resampled.nii.gz
Output Properties
- Dimensions: Match
desired_volumeexactly - Voxel spacing: Automatically adjusted to preserve field of view
- Origin: Preserved from original volume
- Direction: Preserved from original volume
- Data type: Preserved from original volume
Field of View Preservation
The function automatically calculates new voxel spacing to maintain physical dimensions:
new_spacing = original_spacing × (original_size / desired_size)
Example:
Original:
Size: 512 × 512 × 300 voxels
Spacing: 0.5 × 0.5 × 1.0 mm
Field of View: 256 × 256 × 300 mm
After resampling to (256, 256, 128):
Size: 256 × 256 × 128 voxels
Spacing: 1.0 × 1.0 × 2.34 mm (automatically calculated)
Field of View: 256 × 256 × 300 mm (preserved!)
This ensures anatomical structures maintain their true physical size.
Interpolation Method
B-spline Interpolation:
- Provides smooth, continuous interpolation
- Higher quality than linear interpolation
- Reduces artifacts and aliasing
- Suitable for medical imaging applications
- Balances quality and computational cost
Advantages over other methods:
- vs Nearest Neighbor: Much smoother, no blocky artifacts
- vs Linear: Smoother gradients, better for visualization
- vs Higher-order: Good balance of quality and speed
Target Volume Selection
Choose desired_volume based on your requirements:
Common Sizes
| Target Size | Voxel Count | Use Case | Computational Cost |
|---|---|---|---|
| (128, 128, 128) | 2.1M | Fast training, prototyping | Low |
| (224, 224, 128) | 6.4M | Standard deep learning | Medium |
| (256, 256, 256) | 16.8M | High quality, detailed analysis | High |
| (512, 512, 256) | 67.1M | Clinical applications, fine details | Very High |
Selection Guidelines
Consider:
- Input size requirements of your neural network
- Available GPU memory for batch processing
- Level of detail needed for your task
- Processing time constraints
- Storage requirements
Anisotropic sizes (e.g., 224×224×128):
- Common in medical imaging due to slice thickness
- Matches typical CT/MRI acquisition patterns
- Reduces computational cost in slice direction
Exceptions
| Exception | Condition |
|---|---|
FileNotFoundError | The input file does not exist |
ValueError | File is not in .nii.gz format |
ValueError | desired_volume does not contain exactly 3 values |
Usage Notes
- Input Format: Only
.nii.gzfiles are accepted - 3D Volumes Required: Input must be a 3D NIfTI image
- Output Directory: Automatically created if it doesn’t exist
- Metadata Preservation: Origin, direction, and data type are maintained
- Memory Usage: Large target volumes may require significant memory
- Processing Time: Scales with output volume size
Examples
Basic Usage
Resample to standard size:
from nidataset.preprocessing import resampling
resampling(
nii_path="scans/patient_001.nii.gz",
output_path="resampled/",
desired_volume=(224, 224, 128)
)
# Output: resampled/patient_001_resampled.nii.gz
With Debug Output
Enable path printing:
resampling(
nii_path="data/scan.nii.gz",
output_path="processed/",
desired_volume=(256, 256, 256),
debug=True
)
# Prints: Resampled image saved at: 'processed/scan_resampled.nii.gz'
Small Size for Fast Processing
Reduce dimensions for quick training:
resampling(
nii_path="training/full_res.nii.gz",
output_path="training/low_res/",
desired_volume=(128, 128, 128),
debug=True
)
# Faster training with reduced resolution
High Resolution for Clinical Use
Maintain detail for diagnostic applications:
resampling(
nii_path="clinical/diagnostic_scan.nii.gz",
output_path="clinical/processed/",
desired_volume=(512, 512, 256),
debug=True
)
# High resolution for accurate diagnosis
Anisotropic Resampling
Match typical medical imaging dimensions:
resampling(
nii_path="ct_scan.nii.gz",
output_path="preprocessed/",
desired_volume=(224, 224, 96), # Different Z resolution
debug=True
)
# Reflects typical slice thickness
Verifying Field of View Preservation
Check that physical dimensions are maintained:
import nibabel as nib
import numpy as np
from nidataset.preprocessing import resampling
# Original volume
original = nib.load("original/scan.nii.gz")
orig_size = np.array(original.shape)
orig_spacing = np.array(original.header.get_zooms()[:3])
orig_fov = orig_size * orig_spacing
print("Original:")
print(f" Size: {orig_size}")
print(f" Spacing: {orig_spacing}")
print(f" FOV: {orig_fov} mm")
# Resample
target_size = (224, 224, 128)
resampling(
nii_path="original/scan.nii.gz",
output_path="resampled/",
desired_volume=target_size,
debug=True
)
# Resampled volume
resampled = nib.load("resampled/scan_resampled.nii.gz")
resamp_size = np.array(resampled.shape)
resamp_spacing = np.array(resampled.header.get_zooms()[:3])
resamp_fov = resamp_size * resamp_spacing
print("\nResampled:")
print(f" Size: {resamp_size}")
print(f" Spacing: {resamp_spacing}")
print(f" FOV: {resamp_fov} mm")
print(f"\nFOV difference: {np.abs(orig_fov - resamp_fov)} mm")
print(f"FOV preserved: {np.allclose(orig_fov, resamp_fov, atol=0.1)}")
Quality Assessment
Evaluate resampling quality:
import nibabel as nib
import numpy as np
from nidataset.preprocessing import resampling
# Resample
resampling(
nii_path="qa/test.nii.gz",
output_path="qa/resampled/",
desired_volume=(224, 224, 128),
debug=True
)
# Load both
original = nib.load("qa/test.nii.gz")
resampled = nib.load("qa/resampled/test_resampled.nii.gz")
orig_data = original.get_fdata()
resamp_data = resampled.get_fdata()
print("\nQuality Assessment:")
# 1. Dimensions
print(f" Target dimensions: (224, 224, 128)")
print(f" Actual dimensions: {resamp_data.shape}")
print(f" Correct: {resamp_data.shape == (224, 224, 128)}")
# 2. Intensity preservation
print(f"\n Original intensity range: [{orig_data.min():.1f}, {orig_data.max():.1f}]")
print(f" Resampled intensity range: [{resamp_data.min():.1f}, {resamp_data.max():.1f}]")
# 3. Mean intensity (should be similar)
orig_mean = orig_data[orig_data > 0].mean()
resamp_mean = resamp_data[resamp_data > 0].mean()
print(f"\n Original mean: {orig_mean:.2f}")
print(f" Resampled mean: {resamp_mean:.2f}")
print(f" Difference: {abs(orig_mean - resamp_mean):.2f} ({abs(orig_mean - resamp_mean)/orig_mean*100:.1f}%)")
# 4. Check for artifacts
if orig_data.min() >= 0 and resamp_data.min() < 0:
print(f"\n ⚠️ Warning: Negative values introduced")
else:
print(f"\n ✓ No unexpected negative values")
Batch Processing Multiple Files
Resample a set of volumes:
import os
from nidataset.preprocessing import resampling
scan_folder = "raw_scans/"
output_folder = "resampled_scans/"
target_size = (224, 224, 128)
for filename in os.listdir(scan_folder):
if filename.endswith('.nii.gz'):
print(f"Processing {filename}...")
resampling(
nii_path=os.path.join(scan_folder, filename),
output_path=output_folder,
desired_volume=target_size,
debug=True
)
print(f"\nProcessed {len(os.listdir(scan_folder))} files")
Comparing Interpolation Quality
Visualize resampling results:
import nibabel as nib
import matplotlib.pyplot as plt
from nidataset.preprocessing import resampling
# Resample
resampling(
nii_path="visualization/scan.nii.gz",
output_path="visualization/resampled/",
desired_volume=(224, 224, 128),
debug=True
)
# Load and compare
original = nib.load("visualization/scan.nii.gz")
resampled = nib.load("visualization/resampled/scan_resampled.nii.gz")
orig_data = original.get_fdata()
resamp_data = resampled.get_fdata()
# Extract middle slices
orig_mid = orig_data.shape[2] // 2
resamp_mid = resamp_data.shape[2] // 2
fig, axes = plt.subplots(1, 2, figsize=(12, 5))
axes[0].imshow(orig_data[:, :, orig_mid], cmap='gray')
axes[0].set_title(f'Original ({orig_data.shape})')
axes[0].axis('off')
axes[1].imshow(resamp_data[:, :, resamp_mid], cmap='gray')
axes[1].set_title(f'Resampled ({resamp_data.shape})')
axes[1].axis('off')
plt.tight_layout()
plt.savefig('resampling_comparison.png', dpi=150)
print("Comparison saved: resampling_comparison.png")
Memory-Efficient Workflow
Calculate memory requirements:
import nibabel as nib
import numpy as np
from nidataset.preprocessing import resampling
def estimate_memory(shape, dtype='float32'):
"""Estimate memory in GB."""
bytes_per_voxel = np.dtype(dtype).itemsize
total_bytes = np.prod(shape) * bytes_per_voxel
return total_bytes / (1024**3)
# Check original
original = nib.load("large_volume.nii.gz")
orig_memory = estimate_memory(original.shape)
print(f"Original memory: {orig_memory:.2f} GB")
# Calculate resampled memory
target_size = (224, 224, 128)
target_memory = estimate_memory(target_size)
print(f"After resampling: {target_memory:.2f} GB")
print(f"Memory reduction: {(1 - target_memory/orig_memory)*100:.1f}%")
# Proceed with resampling
if target_memory < 2.0: # Less than 2GB
resampling(
nii_path="large_volume.nii.gz",
output_path="reduced/",
desired_volume=target_size,
debug=True
)
print("Resampling completed")
else:
print("Warning: Target size still requires significant memory")
Integration with Deep Learning Pipeline
Prepare data for model input:
from nidataset.preprocessing import resampling
import nibabel as nib
import numpy as np
def prepare_for_training(input_path, output_path, target_size=(224, 224, 128)):
"""Resample and normalize for training."""
# Step 1: Resample
resampling(
nii_path=input_path,
output_path=output_path,
desired_volume=target_size,
debug=True
)
# Step 2: Load resampled
scan_name = os.path.basename(input_path).replace('.nii.gz', '')
resampled_path = f"{output_path}/{scan_name}_resampled.nii.gz"
img = nib.load(resampled_path)
data = img.get_fdata()
# Step 3: Normalize
data = (data - data.mean()) / data.std()
# Step 4: Save normalized
normalized = nib.Nifti1Image(data, img.affine)
normalized_path = f"{output_path}/{scan_name}_ready.nii.gz"
nib.save(normalized, normalized_path)
print(f"Training-ready volume: {normalized_path}")
return normalized_path
# Use in pipeline
prepare_for_training(
"raw_data/scan_001.nii.gz",
"training_ready/",
target_size=(224, 224, 128)
)
Multi-Resolution Analysis
Create pyramid of resolutions:
from nidataset.preprocessing import resampling
input_file = "analysis/scan.nii.gz"
resolutions = {
'coarse': (112, 112, 64),
'medium': (224, 224, 128),
'fine': (448, 448, 256)
}
for level, size in resolutions.items():
print(f"\nCreating {level} resolution...")
resampling(
nii_path=input_file,
output_path=f"multi_res/{level}/",
desired_volume=size,
debug=True
)
print("\nMulti-resolution pyramid created")
Preserving Metadata
Verify all metadata is maintained:
import nibabel as nib
import numpy as np
from nidataset.preprocessing import resampling
# Original
original = nib.load("metadata_test/scan.nii.gz")
orig_affine = original.affine
orig_origin = original.affine[:3, 3]
orig_direction = original.affine[:3, :3]
print("Original metadata:")
print(f" Origin: {orig_origin}")
print(f" Direction:\n{orig_direction}")
# Resample
resampling(
nii_path="metadata_test/scan.nii.gz",
output_path="metadata_test/output/",
desired_volume=(224, 224, 128),
debug=True
)
# Check resampled
resampled = nib.load("metadata_test/output/scan_resampled.nii.gz")
resamp_affine = resampled.affine
resamp_origin = resampled.affine[:3, 3]
resamp_direction = resampled.affine[:3, :3] / np.array(resampled.header.get_zooms()[:3])
print("\nResampled metadata:")
print(f" Origin: {resamp_origin}")
print(f" Direction:\n{resamp_direction}")
print("\nMetadata preserved:")
print(f" Origin match: {np.allclose(orig_origin, resamp_origin)}")
print(f" Direction match: {np.allclose(orig_direction/np.linalg.norm(orig_direction), resamp_direction/np.linalg.norm(resamp_direction))}")
Typical Workflow
from nidataset.preprocessing import resampling
import nibabel as nib
# 1. Check original dimensions
original = nib.load("data/scan.nii.gz")
print(f"Original shape: {original.shape}")
print(f"Original spacing: {original.header.get_zooms()[:3]}")
# 2. Resample to target size
target_size = (224, 224, 128)
resampling(
nii_path="data/scan.nii.gz",
output_path="data/resampled/",
desired_volume=target_size,
debug=True
)
# 3. Verify result
resampled = nib.load("data/resampled/scan_resampled.nii.gz")
print(f"\nResampled shape: {resampled.shape}")
print(f"Resampled spacing: {resampled.header.get_zooms()[:3]}")
# 4. Use resampled volume for:
# - Training neural networks with fixed input size
# - Batch processing without dimension mismatches
# - Reducing computational requirements
# - Standardizing datasets from different sources