mip
Generate a sliding-window Maximum Intensity Projection from a 3D medical imaging volume to enhance high-intensity structures like blood vessels.
mip(
nii_path: str,
output_path: str,
window_size: int = 10,
view: str = "axial",
debug: bool = False
) -> None
Overview
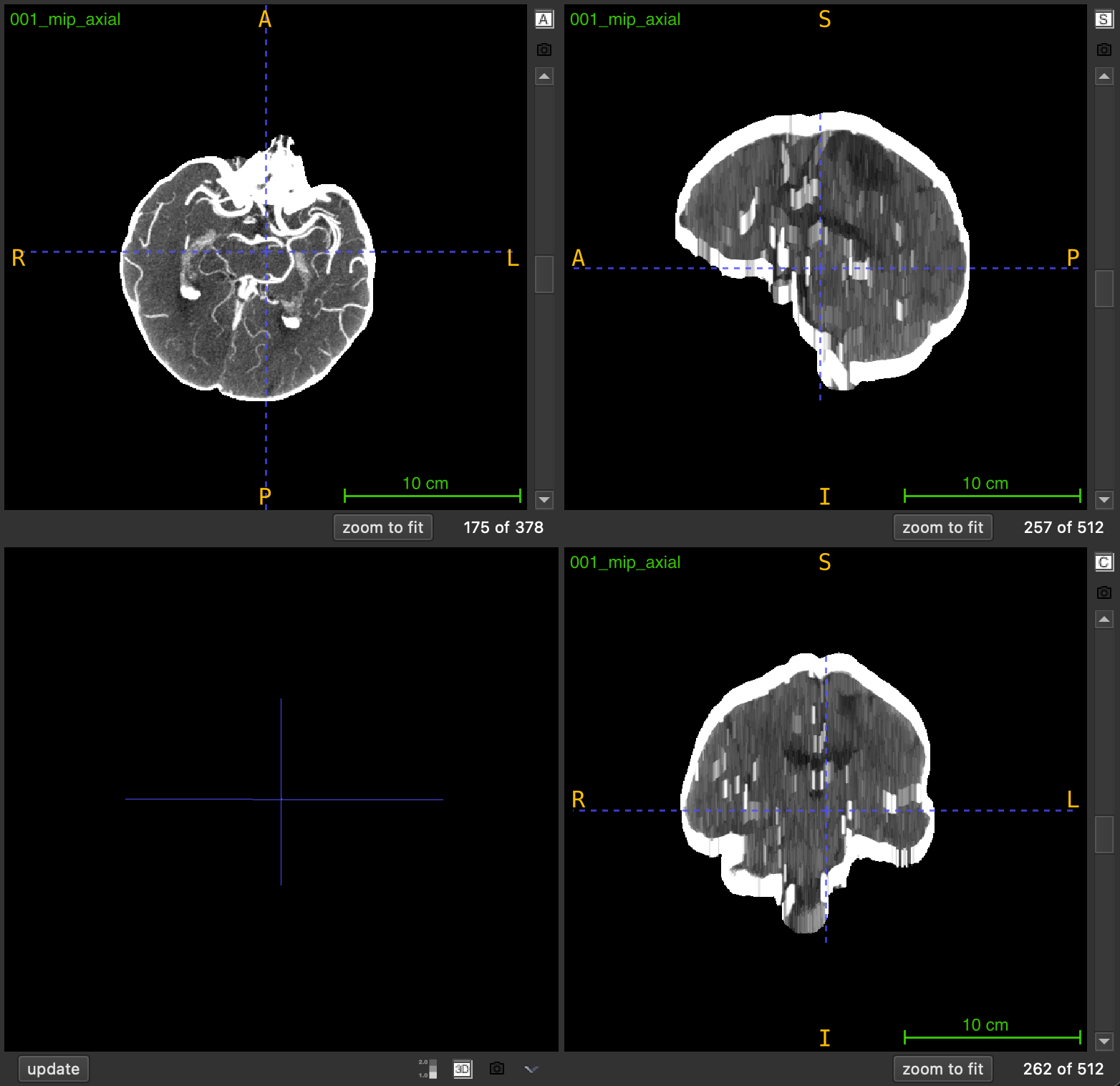
Maximum Intensity Projection (MIP) is an enhancement technique that highlights high-intensity structures by replacing each voxel with the maximum intensity value found in its local neighborhood. This function applies a sliding-window approach where each output slice is created by projecting a local neighborhood of slices along a chosen anatomical axis.
Key characteristics:
- Local projection: Each slice uses a neighborhood centered at its position (not global)
- Same dimensions: Output volume has identical shape to input
- Enhanced visibility: Bright structures (vessels, contrast) become more prominent
- Preserved spatial information: Unlike global MIP, spatial relationships are maintained
The technique is particularly valuable for:
- Enhancing vascular structures in angiography
- Improving contrast-enhanced region visibility
- Preprocessing for vessel detection and segmentation
- Reducing noise while preserving peak intensities
Parameters
| Name | Type | Default | Description |
|---|---|---|---|
nii_path | str | required | Path to the input medical volume in .nii.gz format. |
output_path | str | required | Directory where the MIP volume will be saved. Created automatically if it doesn’t exist. |
window_size | int | 10 | Number of slices on each side of current position for projection. Effective window: 2×size+1. |
view | str | "axial" | Anatomical plane for projection: "axial", "coronal", or "sagittal". |
debug | bool | False | If True, prints progress information and output filename. |
Returns
None – The function saves the MIP volume to disk.
Output File
The MIP volume is saved as:
<PREFIX>_mip_<VIEW>.nii.gz
Example: Input scan_042.nii.gz → Output scan_042_mip_axial.nii.gz
Output Volume Properties
- Dimensions: Same as input volume
- Data type: Same as input (typically float)
- Spatial metadata: Inherits affine transformation from input
- Intensity range: Equal to or higher than input (due to max operation)
Sliding Window Mechanism
For each slice at position i, the function:
- Defines a neighborhood:
[i - window_size, i + window_size] - Extracts the subvolume within this range
- Computes the maximum intensity across the neighborhood
- Assigns the result to output slice
i
Effective window length: 2 × window_size + 1
Example (window_size=3):
Slice index: 10
Window range: [7, 13] (7 slices total)
Output slice 10 = max(slices 7, 8, 9, 10, 11, 12, 13)
Window Size Selection
The window_size parameter controls the neighborhood extent:
| Window Size | Effective Slices | Effect | Best For |
|---|---|---|---|
| 3-5 | 7-11 slices | Minimal smoothing, local enhancement | Fine vessel details |
| 10-15 | 21-31 slices | Moderate enhancement, balanced | General purpose |
| 20-30 | 41-61 slices | Strong enhancement, wider context | Large vessel structures |
| 40+ | 81+ slices | Maximum enhancement, near-global view | Overview visualization |
Trade-off considerations:
- Smaller windows: Preserve fine details but provide less enhancement
- Larger windows: Stronger enhancement but may blur details and mix unrelated structures
- Optimal size: Depends on structure size and spacing between slices
Anatomical Views
The view parameter determines the projection axis:
| View | Projection Axis | Direction | Common Applications |
|---|---|---|---|
"axial" | Z-axis | Top to bottom | Brain vessels, chest imaging |
"coronal" | Y-axis | Front to back | Spine vessels, full body |
"sagittal" | X-axis | Left to right | Bilateral vessel comparison |
Exceptions
| Exception | Condition |
|---|---|
FileNotFoundError | The input file does not exist |
ValueError | File is not in .nii.gz format |
ValueError | Input is not a 3D volume |
ValueError | Invalid view parameter |
Usage Notes
- Input Format: Only
.nii.gzfiles are accepted - 3D Volumes Required: Input must be a 3D NIfTI image
- Edge Handling: Windows are clipped at volume boundaries
- Output Directory: Automatically created if it doesn’t exist
- Spatial Metadata: Affine transformation is preserved
- Progress Display: Shows progress bar during processing
Examples
Basic Usage
Generate MIP with default settings:
from nidataset.preprocessing import mip
mip(
nii_path="angiography/vessel_scan.nii.gz",
output_path="enhanced/",
window_size=10,
view="axial"
)
# Output: enhanced/vessel_scan_mip_axial.nii.gz
With Debug Information
Enable verbose output:
mip(
nii_path="ct_angio.nii.gz",
output_path="mip_results/",
window_size=15,
view="coronal",
debug=True
)
# Prints progress bar and:
# MIP saved at: mip_results/ct_angio_mip_coronal.nii.gz
Strong Enhancement for Large Vessels
Use large window for prominent structures:
mip(
nii_path="aorta_scan.nii.gz",
output_path="enhanced_vessels/",
window_size=25, # 51-slice window
view="axial",
debug=True
)
Fine Detail Preservation
Use small window for detailed structures:
mip(
nii_path="cerebral_vessels.nii.gz",
output_path="detail_preserved/",
window_size=5, # 11-slice window
view="axial",
debug=True
)
Multi-View Processing
Generate MIP for all anatomical views:
from nidataset.preprocessing import mip
scan_file = "vessel_scan.nii.gz"
views = ["axial", "coronal", "sagittal"]
for view in views:
mip(
nii_path=scan_file,
output_path=f"mip_multiview/{view}/",
window_size=15,
view=view,
debug=True
)
# Creates: mip_multiview/axial/vessel_scan_mip_axial.nii.gz, etc.
Comparing Window Sizes
Test different window sizes to find optimal enhancement:
from nidataset.preprocessing import mip
scan_file = "test_scan.nii.gz"
window_sizes = [5, 10, 15, 20, 30]
for size in window_sizes:
mip(
nii_path=scan_file,
output_path=f"window_test/size_{size}/",
window_size=size,
view="axial",
debug=True
)
print(f"Window size {size}: effective window = {2*size+1} slices")
print("\nCompare outputs visually to select optimal window")
Quality Assessment
Generate MIP and evaluate enhancement:
import nibabel as nib
import numpy as np
from nidataset.preprocessing import mip
# Generate MIP
mip(
nii_path="qa/original_scan.nii.gz",
output_path="qa/mip_output/",
window_size=15,
view="axial",
debug=True
)
# Load original and MIP
original = nib.load("qa/original_scan.nii.gz")
mip_vol = nib.load("qa/mip_output/original_scan_mip_axial.nii.gz")
orig_data = original.get_fdata()
mip_data = mip_vol.get_fdata()
# Analyze enhancement
print(f"\nEnhancement Analysis:")
print(f" Original shape: {orig_data.shape}")
print(f" MIP shape: {mip_data.shape}")
print(f" Original intensity range: [{orig_data.min():.1f}, {orig_data.max():.1f}]")
print(f" MIP intensity range: [{mip_data.min():.1f}, {mip_data.max():.1f}]")
# Compare mean intensities in vessel region (example coordinates)
vessel_region_orig = orig_data[100:150, 100:150, 50:100]
vessel_region_mip = mip_data[100:150, 100:150, 50:100]
print(f" Vessel region - Original mean: {vessel_region_orig.mean():.1f}")
print(f" Vessel region - MIP mean: {vessel_region_mip.mean():.1f}")
print(f" Enhancement factor: {vessel_region_mip.mean() / vessel_region_orig.mean():.2f}x")
Preprocessing for Vessel Segmentation
Use MIP as preprocessing step:
from nidataset.preprocessing import mip
from nidataset.volume import generate_brain_mask
scan_file = "angiography.nii.gz"
# Step 1: Generate MIP to enhance vessels
mip(
nii_path=scan_file,
output_path="pipeline/mip/",
window_size=15,
view="axial",
debug=True
)
# Step 2: Generate brain mask on enhanced volume
generate_brain_mask(
nii_path="pipeline/mip/angiography_mip_axial.nii.gz",
output_path="pipeline/masks/",
threshold=(100, 500), # Higher threshold for enhanced vessels
closing_radius=3,
debug=True
)
# Step 3: Apply mask to MIP volume
import nibabel as nib
mip_vol = nib.load("pipeline/mip/angiography_mip_axial.nii.gz")
mask = nib.load("pipeline/masks/angiography_mip_axial_brain_mask.nii.gz")
mip_data = mip_vol.get_fdata()
mask_data = mask.get_fdata()
vessel_focused = mip_data * mask_data
# Save vessel-focused volume
vessel_img = nib.Nifti1Image(vessel_focused, mip_vol.affine)
nib.save(vessel_img, "pipeline/vessel_segmentation_ready.nii.gz")
print("Preprocessing complete - ready for vessel segmentation")
Optimal Window Size Determination
Analyze enhancement vs detail trade-off:
import nibabel as nib
import numpy as np
from nidataset.preprocessing import mip
from scipy import ndimage
scan_file = "analysis/test_scan.nii.gz"
window_sizes = [5, 10, 15, 20, 25, 30]
results = []
for size in window_sizes:
# Generate MIP
mip(
nii_path=scan_file,
output_path=f"analysis/window_{size}/",
window_size=size,
view="axial"
)
# Load and analyze
original = nib.load(scan_file)
mip_vol = nib.load(f"analysis/window_{size}/test_scan_mip_axial.nii.gz")
orig_data = original.get_fdata()
mip_data = mip_vol.get_fdata()
# Calculate metrics
enhancement = mip_data.mean() / orig_data.mean()
# Estimate detail preservation (via edge detection)
edges_orig = ndimage.sobel(orig_data)
edges_mip = ndimage.sobel(mip_data)
edge_ratio = np.sum(edges_mip > 0) / np.sum(edges_orig > 0)
results.append({
'window_size': size,
'effective_slices': 2*size+1,
'enhancement': enhancement,
'edge_preservation': edge_ratio
})
# Display results
import pandas as pd
df = pd.DataFrame(results)
print("\nWindow Size Analysis:")
print(df.to_string(index=False))
print("\nOptimal: Balance between enhancement and edge preservation")
Creating Visualization Comparison
Generate side-by-side comparison:
import nibabel as nib
import numpy as np
from nidataset.preprocessing import mip
# Generate MIP
mip(
nii_path="visualization/scan.nii.gz",
output_path="visualization/mip/",
window_size=15,
view="axial",
debug=True
)
# Load both volumes
original = nib.load("visualization/scan.nii.gz")
mip_vol = nib.load("visualization/mip/scan_mip_axial.nii.gz")
orig_data = original.get_fdata()
mip_data = mip_vol.get_fdata()
# Create side-by-side comparison (middle slice)
mid_slice = orig_data.shape[2] // 2
orig_slice = orig_data[:, :, mid_slice]
mip_slice = mip_data[:, :, mid_slice]
# Normalize for display
orig_norm = (orig_slice - orig_slice.min()) / (orig_slice.max() - orig_slice.min())
mip_norm = (mip_slice - mip_slice.min()) / (mip_slice.max() - mip_slice.min())
# Concatenate horizontally
comparison = np.hstack([orig_norm, mip_norm])
# Save as 2D image for visualization
from PIL import Image
comparison_img = Image.fromarray((comparison * 255).astype(np.uint8))
comparison_img.save("visualization/comparison.png")
print("Comparison image created: visualization/comparison.png")
Integration with Deep Learning Pipeline
Prepare MIP-enhanced data for training:
from nidataset.preprocessing import mip
from nidataset.slices import extract_slices
scan_files = ["train_001.nii.gz", "train_002.nii.gz", "train_003.nii.gz"]
for scan_file in scan_files:
# Step 1: Generate MIP
mip(
nii_path=f"training/raw/{scan_file}",
output_path="training/mip_volumes/",
window_size=15,
view="axial",
debug=True
)
# Step 2: Extract 2D slices from MIP
mip_filename = scan_file.replace('.nii.gz', '_mip_axial.nii.gz')
extract_slices(
nii_path=f"training/mip_volumes/{mip_filename}",
output_path="training/mip_slices/",
view="axial",
target_size=(512, 512),
debug=True
)
print("MIP-enhanced training data prepared")
Batch Processing with Error Handling
Process multiple files robustly:
from nidataset.preprocessing import mip
import os
scan_folder = "raw_scans/"
output_folder = "mip_processed/"
failed_files = []
for filename in os.listdir(scan_folder):
if filename.endswith('.nii.gz'):
try:
mip(
nii_path=os.path.join(scan_folder, filename),
output_path=output_folder,
window_size=15,
view="axial",
debug=True
)
print(f"✓ Processed: {filename}")
except Exception as e:
print(f"✗ Failed: {filename} - {str(e)}")
failed_files.append(filename)
if failed_files:
print(f"\n{len(failed_files)} files failed:")
for f in failed_files:
print(f" - {f}")
else:
print("\nAll files processed successfully")
Typical Workflow
from nidataset.preprocessing import mip
import nibabel as nib
# 1. Define input and output
scan_file = "angiography/vessel_scan.nii.gz"
output_folder = "enhanced/"
# 2. Generate MIP with appropriate window size
mip(
nii_path=scan_file,
output_path=output_folder,
window_size=15, # Adjust based on vessel size
view="axial",
debug=True
)
# 3. Verify output
mip_vol = nib.load("enhanced/vessel_scan_mip_axial.nii.gz")
print(f"MIP volume shape: {mip_vol.shape}")
# 4. Use MIP volume for:
# - Vessel detection and segmentation
# - Aneurysm identification
# - Vascular analysis
# - Training data for deep learning models