crop_and_pad
Crop a medical volume to its non-zero bounding box, then pad or crop it to a uniform target shape while preserving spatial orientation.
crop_and_pad(
nii_path: str,
output_path: str,
target_shape: tuple = (128, 128, 128),
debug: bool = False
) -> None
Overview
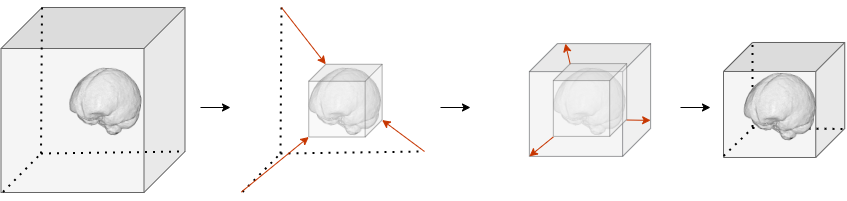
This function standardizes medical imaging volumes by removing empty space and ensuring consistent dimensions. The processing follows these steps:
- Crop to bounding box: Identifies and removes empty voxels around the non-zero region
- Resize to target: Applies symmetric padding if the volume is smaller than the target shape, or centered cropping if it’s larger
- Preserve orientation: Maintains spatial alignment by updating the affine transformation matrix
The output is saved with the suffix _cropped_padded.nii.gz.
Parameters
| Name | Type | Default | Description |
|---|---|---|---|
nii_path | str | required | Path to the input volume in .nii.gz format. |
output_path | str | required | Directory where the processed scan will be saved. Created automatically if it doesn’t exist. |
target_shape | tuple | (128, 128, 128) | Desired output shape (X, Y, Z) after cropping and padding. |
debug | bool | False | If True, prints detailed shape information and output file path to console. |
Returns
None – The function saves the processed volume to disk.
Output File
The processed volume is saved as:
<PREFIX>_cropped_padded.nii.gz
where <PREFIX> is the original filename without the .nii.gz extension.
Example: Input case_01.nii.gz → Output case_01_cropped_padded.nii.gz
Exceptions
| Exception | Condition |
|---|---|
FileNotFoundError | The input file does not exist |
ValueError | The file is not in .nii.gz format |
ValueError | The input file is not 3D (wrong number of dimensions) |
ValueError | The target shape is not 3D |
Processing Details
Cropping Strategy
When the cropped volume exceeds the target shape in any dimension, centered cropping is applied to that dimension. This preserves the central portion of the scan.
Padding Strategy
When the cropped volume is smaller than the target shape in any dimension, symmetric padding with zero values is applied. If the required padding is odd, the extra voxel is placed on the right/bottom/back side.
Spatial Alignment
The affine transformation matrix and NIfTI header are automatically updated to maintain correct spatial orientation and world coordinates. This ensures that the processed volume aligns correctly with the original anatomical space.
Usage Notes
- Input Format: Only
.nii.gzfiles are accepted - 3D Volumes Required: Input must be a 3D NIfTI image
- Output Directory: Automatically created if it doesn’t exist
- Affine Preservation: The spatial transformation matrix is adjusted to maintain anatomical alignment
- Header Preservation: Original header metadata is retained where possible
Examples
Basic Usage
Process a single volume with default 128³ output:
from nidataset.volume import crop_and_pad
crop_and_pad(
nii_path="dataset/scan_001.nii.gz",
output_path="preprocessed/",
target_shape=(128, 128, 128)
)
# Output: preprocessed/scan_001_cropped_padded.nii.gz
With Debug Information
Enable detailed output to verify processing:
crop_and_pad(
nii_path="dataset/scan_001.nii.gz",
output_path="preprocessed/",
target_shape=(128, 128, 128),
debug=True
)
# Prints:
# Input File: 'dataset/scan_001.nii.gz'
# Output Path: 'preprocessed/'
# Original Shape: (512, 512, 300) | Cropped Shape: (420, 380, 250) | Final Shape: (128, 128, 128)
# Processed volume saved at: preprocessed/scan_001_cropped_padded.nii.gz
Custom Target Shape
Use non-cubic dimensions for specialized applications:
crop_and_pad(
nii_path="data/patient_042.nii.gz",
output_path="output/standardized/",
target_shape=(160, 160, 96)
)
Processing Multiple Files
Combine with a loop for batch processing:
import os
from nidataset.volume import crop_and_pad
input_folder = "raw_scans/"
output_folder = "processed_scans/"
for filename in os.listdir(input_folder):
if filename.endswith(".nii.gz"):
crop_and_pad(
nii_path=os.path.join(input_folder, filename),
output_path=output_folder,
target_shape=(128, 128, 128),
debug=True
)
Verifying Output Shape
Check that processing produced the expected dimensions:
import nibabel as nib
from nidataset.volume import crop_and_pad
# Process the volume
crop_and_pad(
nii_path="input/scan.nii.gz",
output_path="output/",
target_shape=(256, 256, 256)
)
# Verify output shape
img = nib.load("output/scan_cropped_padded.nii.gz")
print(f"Output shape: {img.shape}") # Should be (256, 256, 256)
Typical Workflow
from nidataset.volume import crop_and_pad
# 1. Define input and output paths
input_scan = "raw_data/patient_scan.nii.gz"
output_dir = "preprocessed_data/"
# 2. Set target dimensions based on your model requirements
target_dims = (128, 128, 128)
# 3. Process the volume
crop_and_pad(
nii_path=input_scan,
output_path=output_dir,
target_shape=target_dims,
debug=True
)
# 4. Use the processed volume for analysis or training
processed_path = "preprocessed_data/patient_scan_cropped_padded.nii.gz"