extract_bounding_boxes
Detect connected components in a segmentation mask, filter by physical volume, and generate a binary mask containing 3D bounding boxes around valid regions.
extract_bounding_boxes(
mask_path: str,
output_path: str,
voxel_size: tuple = (3.0, 3.0, 3.0),
volume_threshold: float = 1000.0,
mask_value: int = 1,
debug: bool = False
) -> None
Overview
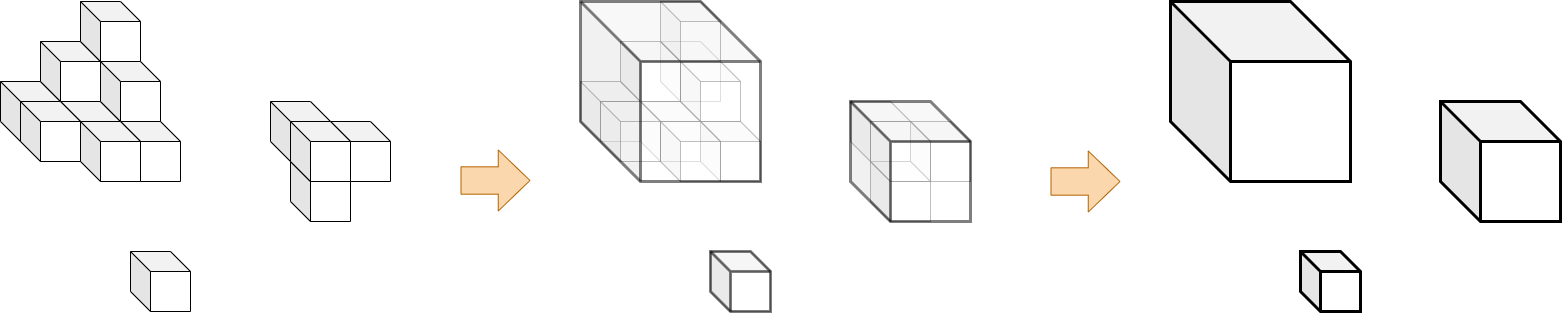
This function converts segmentation masks into simplified bounding box representations by identifying connected components and drawing 3D boxes around structures that meet a minimum volume threshold. The process:
- Identifies connected components in the mask using the specified label value
- Calculates physical volumes in cubic millimeters for each component
- Filters components based on the volume threshold to remove noise or artifacts
- Generates bounding boxes as filled 3D rectangles in a new binary mask
The output is useful for:
- Visualization and quality control of segmentations
- Creating simplified region representations
- Removing small false positives from automated segmentations
- Generating region proposals for detection tasks
Parameters
| Name | Type | Default | Description |
|---|---|---|---|
mask_path | str | required | Path to the input segmentation mask in .nii.gz format. |
output_path | str | required | Directory where the bounding box mask will be saved. Created automatically if it doesn’t exist. |
voxel_size | tuple | (3.0, 3.0, 3.0) | Physical voxel dimensions in millimeters as (x, y, z) for volume calculations. |
volume_threshold | float | 1000.0 | Minimum component volume in mm³ required to generate a bounding box. |
mask_value | int | 1 | Integer label value in the mask representing the target region to analyze. |
debug | bool | False | If True, prints detailed information about detected components and output path. |
Returns
None – The function saves the bounding box mask to disk.
Output File
The function creates a binary mask file:
<PREFIX>_bounding_boxes.nii.gz
where <PREFIX> is the original filename without the .nii.gz extension.
Example: Input lesion_mask.nii.gz → Output lesion_mask_bounding_boxes.nii.gz
Output Mask Properties
- Data type: uint8 (8-bit unsigned integer)
- Box voxels: Value
255(white in most viewers) - Background: Value
0(black) - Spatial metadata: Inherits affine transformation from input mask
Volume Filtering
The function filters connected components based on their physical volume:
Volume Calculation:
volume (mm³) = number_of_voxels × voxel_x × voxel_y × voxel_z
Filtering Logic:
- Components with
volume ≥ volume_threshold→ Bounding box created - Components with
volume < volume_threshold→ Excluded from output
This filtering is essential for:
- Removing imaging noise and artifacts
- Excluding small false positives from automated segmentations
- Focusing on clinically significant structures
- Reducing computational load in downstream processing
Connected Component Analysis
Each disconnected region with the specified mask_value is treated as a separate component:
- Multiple lesions: Each lesion gets its own bounding box
- Single structure: One connected structure produces one box
- Empty mask: No components result in an empty output mask
Bounding boxes are the minimal axis-aligned rectangles that completely contain each component.
Exceptions
| Exception | Condition |
|---|---|
FileNotFoundError | The input mask file does not exist |
ValueError | File is not in .nii.gz format |
ValueError | Input is not a 3D NIfTI image |
Usage Notes
- Input Format: Only
.nii.gzfiles are accepted - 3D Volumes Required: Input must be a 3D NIfTI mask
- Output Directory: Automatically created if it doesn’t exist
- Voxel Size Accuracy: Ensure
voxel_sizematches your scan’s actual resolution for accurate volume filtering - Progress Display: Shows progress bar during component processing
- Box Representation: Bounding boxes are filled volumes, not just edges
Examples
Basic Usage
Extract bounding boxes with default settings:
from nidataset.volume import extract_bounding_boxes
extract_bounding_boxes(
mask_path="segmentations/tumor_mask.nii.gz",
output_path="bounding_boxes/",
voxel_size=(3.0, 3.0, 3.0),
volume_threshold=1000.0,
mask_value=1
)
# Output: bounding_boxes/tumor_mask_bounding_boxes.nii.gz
High-Resolution Scans
Adjust parameters for high-resolution imaging:
extract_bounding_boxes(
mask_path="hr_scans/lesion_segmentation.nii.gz",
output_path="hr_bboxes/",
voxel_size=(0.5, 0.5, 1.0), # High-resolution voxel spacing
volume_threshold=200.0, # Lower threshold for smaller voxels
mask_value=1,
debug=True
)
# Debug output shows number of boxes extracted
Aggressive Noise Filtering
Remove small artifacts with high volume threshold:
extract_bounding_boxes(
mask_path="noisy_predictions/model_output.nii.gz",
output_path="filtered_boxes/",
voxel_size=(2.0, 2.0, 2.0),
volume_threshold=5000.0, # Keep only large structures
mask_value=1,
debug=True
)
print("Small false positives filtered out")
Processing Different Structures
Extract boxes for specific anatomical labels:
from nidataset.volume import extract_bounding_boxes
# Multi-label mask with different organs
mask_file = "multi_label_segmentation.nii.gz"
voxel_dims = (1.0, 1.0, 1.5)
structures = {
'liver': {'value': 1, 'threshold': 50000.0},
'kidneys': {'value': 2, 'threshold': 15000.0},
'spleen': {'value': 3, 'threshold': 8000.0}
}
for name, params in structures.items():
extract_bounding_boxes(
mask_path=mask_file,
output_path=f"structures/{name}/",
voxel_size=voxel_dims,
volume_threshold=params['threshold'],
mask_value=params['value'],
debug=True
)
Quality Control Workflow
Verify segmentation results by examining bounding boxes:
import nibabel as nib
from nidataset.volume import extract_bounding_boxes
# Extract bounding boxes
extract_bounding_boxes(
mask_path="qa/test_segmentation.nii.gz",
output_path="qa/boxes/",
voxel_size=(1.0, 1.0, 1.0),
volume_threshold=1000.0,
mask_value=1,
debug=True
)
# Load and verify output
bbox_img = nib.load("qa/boxes/test_segmentation_bounding_boxes.nii.gz")
bbox_data = bbox_img.get_fdata()
print(f"\nBounding Box Mask Statistics:")
print(f" Shape: {bbox_data.shape}")
print(f" Non-zero voxels: {np.count_nonzero(bbox_data)}")
print(f" Unique values: {np.unique(bbox_data)}")
# Load in viewer for visual inspection
# Open both original mask and bounding box mask in ITK-SNAP or 3D Slicer
Comparing Thresholds
Test different volume thresholds to find optimal filtering:
import nibabel as nib
from nidataset.volume import extract_bounding_boxes
mask_file = "test_mask.nii.gz"
thresholds = [500.0, 1000.0, 2000.0, 5000.0]
for threshold in thresholds:
output_path = f"threshold_test/{int(threshold)}mm3/"
extract_bounding_boxes(
mask_path=mask_file,
output_path=output_path,
voxel_size=(1.0, 1.0, 1.0),
volume_threshold=threshold,
mask_value=1,
debug=True
)
# Count boxes in output
bbox_img = nib.load(f"{output_path}/test_mask_bounding_boxes.nii.gz")
bbox_data = bbox_img.get_fdata()
num_voxels = np.count_nonzero(bbox_data)
print(f"Threshold {threshold} mm³: {num_voxels} voxels in boxes")
print("\nCompare outputs visually to choose optimal threshold")
Creating Region Proposals
Generate simplified region representations for detection models:
from nidataset.volume import extract_bounding_boxes
# Original segmentation from automated method
extract_bounding_boxes(
mask_path="predictions/automated_segmentation.nii.gz",
output_path="region_proposals/",
voxel_size=(1.5, 1.5, 2.0),
volume_threshold=2000.0,
mask_value=1,
debug=True
)
# Use bounding boxes as region proposals for:
# - Fine-tuning detection models
# - Focused segmentation refinement
# - Attention mechanisms
Batch Processing with Different Settings
Process multiple masks with varying parameters:
import os
from nidataset.volume import extract_bounding_boxes
masks = {
'lesions/patient_001.nii.gz': {'threshold': 500.0, 'value': 1},
'lesions/patient_002.nii.gz': {'threshold': 800.0, 'value': 1},
'organs/liver_mask.nii.gz': {'threshold': 50000.0, 'value': 2}
}
voxel_dims = (1.0, 1.0, 1.5)
for mask_path, params in masks.items():
if os.path.exists(mask_path):
case_name = os.path.basename(mask_path).replace('.nii.gz', '')
extract_bounding_boxes(
mask_path=mask_path,
output_path=f"processed_boxes/{case_name}/",
voxel_size=voxel_dims,
volume_threshold=params['threshold'],
mask_value=params['value'],
debug=True
)
Visualizing Bounding Boxes
Create overlays for visual inspection:
import nibabel as nib
import numpy as np
from nidataset.volume import extract_bounding_boxes
# Extract bounding boxes
extract_bounding_boxes(
mask_path="masks/lesion_mask.nii.gz",
output_path="visualization/",
voxel_size=(1.0, 1.0, 1.0),
volume_threshold=1000.0,
mask_value=1
)
# Load original scan and bounding boxes
scan = nib.load("scans/original_scan.nii.gz")
scan_data = scan.get_fdata()
bbox = nib.load("visualization/lesion_mask_bounding_boxes.nii.gz")
bbox_data = bbox.get_fdata()
# Create overlay (boxes have edges highlighted)
overlay = scan_data.copy()
overlay[bbox_data > 0] = scan_data.max() # Highlight box regions
# Save overlay for visualization
overlay_img = nib.Nifti1Image(overlay, scan.affine)
nib.save(overlay_img, "visualization/overlay.nii.gz")
print("Overlay created: visualization/overlay.nii.gz")
Integration with Segmentation Pipeline
Use bounding boxes for validation and refinement:
import nibabel as nib
from nidataset.volume import extract_bounding_boxes
# Step 1: Run automated segmentation (example output)
segmentation_mask = "pipeline/automated_seg.nii.gz"
# Step 2: Extract bounding boxes with filtering
extract_bounding_boxes(
mask_path=segmentation_mask,
output_path="pipeline/validation/",
voxel_size=(1.5, 1.5, 2.0),
volume_threshold=1500.0,
mask_value=1,
debug=True
)
# Step 3: Compare with ground truth
ground_truth = nib.load("pipeline/ground_truth.nii.gz")
gt_data = ground_truth.get_fdata()
bbox_result = nib.load("pipeline/validation/automated_seg_bounding_boxes.nii.gz")
bbox_data = bbox_result.get_fdata()
# Calculate overlap metrics
intersection = np.logical_and(gt_data > 0, bbox_data > 0).sum()
union = np.logical_or(gt_data > 0, bbox_data > 0).sum()
iou = intersection / union if union > 0 else 0
print(f"Bounding box IoU with ground truth: {iou:.3f}")
Analyzing Component Sizes
Extract boxes and analyze size distribution:
import nibabel as nib
import numpy as np
from scipy import ndimage as ndi
from nidataset.volume import extract_bounding_boxes
# Extract bounding boxes
extract_bounding_boxes(
mask_path="masks/multi_lesion.nii.gz",
output_path="analysis/",
voxel_size=(1.0, 1.0, 1.0),
volume_threshold=500.0,
mask_value=1,
debug=True
)
# Load original mask to analyze components
mask_img = nib.load("masks/multi_lesion.nii.gz")
mask_data = mask_img.get_fdata()
binary_mask = (mask_data == 1).astype(np.uint8)
# Get component labels
labeled, num_components = ndi.label(binary_mask)
# Calculate volumes
volumes = []
for label in range(1, num_components + 1):
volume_mm3 = np.sum(labeled == label) * 1.0 * 1.0 * 1.0
if volume_mm3 >= 500.0: # Match threshold
volumes.append(volume_mm3)
print(f"\nComponent Volume Analysis:")
print(f" Total components: {num_components}")
print(f" Components above threshold: {len(volumes)}")
print(f" Volume range: {min(volumes):.1f} - {max(volumes):.1f} mm³")
print(f" Average volume: {np.mean(volumes):.1f} mm³")
Typical Workflow
from nidataset.volume import extract_bounding_boxes
import nibabel as nib
# 1. Define input parameters
mask_file = "segmentations/patient_042_lesions.nii.gz"
output_folder = "bounding_boxes/"
voxel_spacing = (1.5, 1.5, 2.0) # Your scan's voxel dimensions
min_volume = 1000.0 # Minimum lesion volume to consider
# 2. Extract bounding boxes
extract_bounding_boxes(
mask_path=mask_file,
output_path=output_folder,
voxel_size=voxel_spacing,
volume_threshold=min_volume,
mask_value=1,
debug=True
)
# 3. Verify output
bbox_file = "bounding_boxes/patient_042_lesions_bounding_boxes.nii.gz"
bbox_img = nib.load(bbox_file)
print(f"Created bounding box mask: {bbox_img.shape}")
# 4. Use bounding boxes for:
# - Visualization in medical imaging viewers
# - Region proposal generation
# - Quality control of segmentations
# - Simplified structure representation