register_mask
Apply a saved transformation to a mask using a reference registered image.
register_mask(
mask_path: str,
transform_path: str,
registered_path: str,
output_path: str,
is_binary: bool = True,
debug: bool = False
) -> None
Overview
This function applies a previously computed transformation (from registration) to align a mask to the same space as a registered image. It’s essential for propagating brain masks, segmentation masks, or ROI masks through registration workflows while maintaining spatial correspondence.
Transformation pipeline:
- Load: Reads mask, transformation, and reference image
- Interpolation selection: Chooses method based on mask type (binary vs. continuous)
- Resampling: Applies transformation with appropriate interpolation
- Output: Saves transformed mask in template space
This is essential for:
- Propagating segmentations to normalized space
- Maintaining label correspondence across registrations
- Applying vessel masks, lesion masks, or ROI masks to registered volumes
- Ensuring spatial consistency in multi-modal analyses
- Creating population-based mask atlases
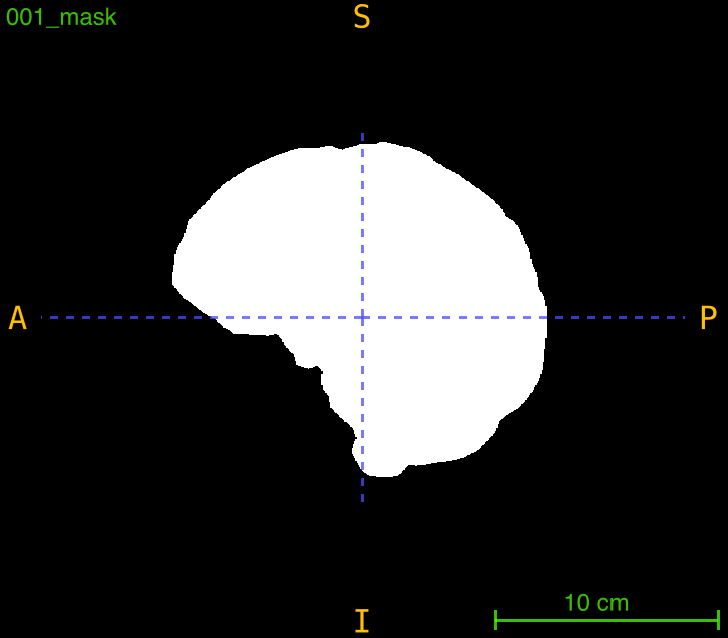
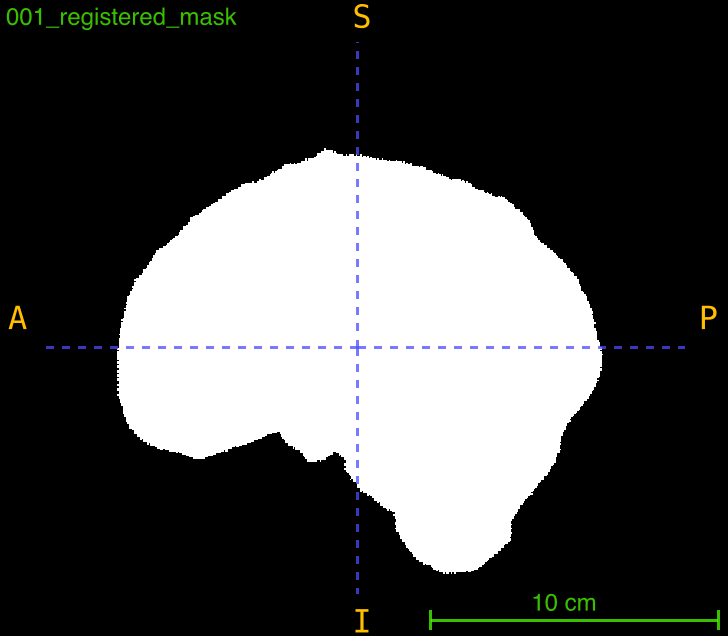
Parameters
| Name | Type | Default | Description |
|---|---|---|---|
mask_path | str | required | Path to the input mask .nii.gz file to be transformed. Must end with _mask.nii.gz. |
transform_path | str | required | Path to the transformation file (.tfm) from a previous registration. |
registered_path | str | required | Path to the registered image that defines the target space and grid. |
output_path | str | required | Path where the transformed mask will be saved (including filename). |
is_binary | bool | True | If True, uses nearest neighbor interpolation. If False, uses linear interpolation. |
debug | bool | False | If True, prints detailed information about the transformation process. |
Returns
None — The function saves the transformed mask to disk.
Output Files
The function generates a single file:
| File | Description | Interpolation Method |
|---|---|---|
<FILENAME>_registered_mask.nii.gz | Mask aligned to template space | Nearest neighbor or linear |
Example: Input lesion_mask.nii.gz produces lesion_registered_mask.nii.gz
Interpolation Methods
The function selects interpolation based on the is_binary parameter:
Nearest Neighbor Interpolation (is_binary=True)
- Use for: Binary masks, label maps, segmentation masks
- Preserves: Discrete label values (0, 1, 2, etc.)
- Advantages: No interpolation artifacts, maintains exact labels
- Examples: Brain masks, vessel segmentations, lesion labels
Linear Interpolation (is_binary=False)
- Use for: Probability maps, continuous-valued masks, partial volume maps
- Creates: Smooth transitions between values
- Advantages: Better for continuous data
- Examples: Tissue probability maps, gradient maps, confidence scores
Interpolation Selection Guide
| Mask Type | is_binary Setting | Reason |
|---|---|---|
| Binary segmentation | True | Preserves discrete labels (0, 1) |
| Multi-label segmentation | True | Preserves label integrity (0, 1, 2, 3, …) |
| Vessel masks | True | Maintains vessel vs. background distinction |
| Lesion masks | True | Preserves lesion boundaries |
| Tissue probability maps | False | Allows smooth probability values |
| Confidence maps | False | Maintains continuous confidence scores |
| Gradient maps | False | Preserves smooth intensity transitions |
| Distance maps | False | Maintains continuous distance values |
Important Notes
Mask Naming Convention
- All mask files must end with
_mask.nii.gz - This is enforced by the function’s validation
- Examples of valid names:
patient001_brain_mask.nii.gz(valid)lesion_mask.nii.gz(valid)vessel_segmentation_mask.nii.gz(valid)
- Examples of invalid names:
patient001_brain.nii.gz(missing_mask)lesion.nii.gz(missing_mask)mask_brain.nii.gz(_masknot at end)
File Matching in Workflows
When using register_mask_dataset, the function will:
- Look for masks ending with
_mask.nii.gz - Match them to transformations by stripping the
_masksuffix - Example:
patient001_brain_mask.nii.gzmatchespatient001_brain_transformation.tfm
Registration Workflow Integration
This function is designed to work seamlessly with register_CTA:
1. register_CTA()
└── Produces: transformation.tfm + registered_image.nii.gz
2. register_mask()
└── Uses: transformation.tfm + registered_image.nii.gz
└── Applies to: any additional mask
Exceptions
| Exception | Condition |
|---|---|
FileNotFoundError | Any required input file does not exist |
ValueError | Input mask does not end with _mask.nii.gz |
Usage Notes
- Input Format: Mask files must end with
_mask.nii.gz(enforced by validation) - 3D Masks Required: Input must be 3D NIfTI image
- Transform Dependency: Transformation must be from a completed registration
- Reference Space: Output matches the space and dimensions of the reference image
- Output Directories: Automatically created if they don’t exist
- Pixel Type Preservation: Output maintains the same pixel type as input mask
Examples
Basic Usage
Apply transformation to a vessel mask:
from nidataset.preprocessing import register_CTA, register_mask
# Step 1: Register the main CTA scan
register_CTA(
nii_path="scan.nii.gz",
mask_path="scan_mask.nii.gz",
template_path="template.nii.gz",
template_mask_path="template_mask.nii.gz",
output_path="registered/",
debug=True
)
# Step 2: Apply the same transformation to vessel mask
register_mask(
mask_path="scan_vessel_mask.nii.gz",
transform_path="registered/scan_transformation.tfm",
registered_path="registered/scan_registered.nii.gz",
output_path="registered/scan_vessel_registered_mask.nii.gz",
is_binary=True,
debug=True
)
# Prints: Registered mask saved at: 'registered/scan_vessel_registered_mask.nii.gz'
Register Multiple Masks
Apply transformation to several masks from the same scan:
from nidataset.preprocessing import register_CTA, register_mask
import os
# Register the main CTA scan
register_CTA(
nii_path="patient001.nii.gz",
mask_path="patient001_brain_mask.nii.gz",
template_path="template.nii.gz",
template_mask_path="template_mask.nii.gz",
output_path="output/patient001/"
)
# List of additional masks to register
masks_to_register = [
"patient001_lesion_mask.nii.gz",
"patient001_vessel_mask.nii.gz",
"patient001_csf_mask.nii.gz",
"patient001_hemorrhage_mask.nii.gz"
]
# Apply transformation to all masks
for mask_file in masks_to_register:
base_name = mask_file.replace("_mask.nii.gz", "")
output_name = f"{base_name}_registered_mask.nii.gz"
register_mask(
mask_path=mask_file,
transform_path="output/patient001/patient001_transformation.tfm",
registered_path="output/patient001/patient001_registered.nii.gz",
output_path=f"output/patient001/{output_name}",
is_binary=True
)
print(f"Registered: {mask_file}")
Register Probability Maps
Apply transformation to continuous-valued masks:
from nidataset.preprocessing import register_mask
# Register a tissue probability map
register_mask(
mask_path="probability_maps/gray_matter_prob_mask.nii.gz",
transform_path="transforms/case001_transformation.tfm",
registered_path="registered/case001_registered.nii.gz",
output_path="registered/gray_matter_prob_registered_mask.nii.gz",
is_binary=False, # Use linear interpolation for smooth values
debug=True
)
# Register white matter probability
register_mask(
mask_path="probability_maps/white_matter_prob_mask.nii.gz",
transform_path="transforms/case001_transformation.tfm",
registered_path="registered/case001_registered.nii.gz",
output_path="registered/white_matter_prob_registered_mask.nii.gz",
is_binary=False,
debug=True
)
Error Handling for Missing Masks
Handle cases where some masks might not exist:
from nidataset.preprocessing import register_mask
import os
cases = ["case001", "case002", "case003"]
mask_types = ["lesion", "vessel", "hemorrhage"]
for case in cases:
for mask_type in mask_types:
mask_path = f"masks/{case}_{mask_type}_mask.nii.gz"
# Check if mask exists before attempting registration
if not os.path.exists(mask_path):
print(f"Skipping {case} - {mask_type}: mask not found")
continue
try:
output_path = f"registered/{case}/{case}_{mask_type}_registered_mask.nii.gz"
register_mask(
mask_path=mask_path,
transform_path=f"registered/{case}/{case}_transformation.tfm",
registered_path=f"registered/{case}/{case}_registered.nii.gz",
output_path=output_path,
is_binary=True
)
print(f"{case} - {mask_type}: registered successfully")
except Exception as e:
print(f"{case} - {mask_type}: failed - {str(e)}")
Multi-Label Mask Registration
Register masks with multiple discrete labels:
from nidataset.preprocessing import register_mask
import nibabel as nib
import numpy as np
# Register a multi-label segmentation mask
register_mask(
mask_path="segmentation/anatomical_labels_mask.nii.gz",
transform_path="transforms/patient_transformation.tfm",
registered_path="registered/patient_registered.nii.gz",
output_path="registered/anatomical_labels_registered_mask.nii.gz",
is_binary=True, # Preserves discrete labels (0, 1, 2, 3, ...)
debug=True
)
# Verify label preservation
original = nib.load("segmentation/anatomical_labels_mask.nii.gz").get_fdata()
registered = nib.load("registered/anatomical_labels_registered_mask.nii.gz").get_fdata()
original_labels = np.unique(original)
registered_labels = np.unique(registered)
print(f"\nLabel Verification:")
print(f" Original labels: {original_labels}")
print(f" Registered labels: {registered_labels}")
print(f" Labels preserved: {set(original_labels) == set(registered_labels)}")
Creating Population-Based Mask Atlas
Combine registered masks from multiple subjects:
from nidataset.preprocessing import register_CTA, register_mask
import nibabel as nib
import numpy as np
# Register scans and masks for all subjects
subjects = ["sub001", "sub002", "sub003", "sub004", "sub005"]
# Step 1: Register all subjects to template
for subject in subjects:
register_CTA(
nii_path=f"data/{subject}.nii.gz",
mask_path=f"data/{subject}_brain_mask.nii.gz",
template_path="template.nii.gz",
template_mask_path="template_mask.nii.gz",
output_path=f"registered/{subject}/",
cleanup=True
)
# Register lesion mask
register_mask(
mask_path=f"data/{subject}_lesion_mask.nii.gz",
transform_path=f"registered/{subject}/{subject}_transformation.tfm",
registered_path=f"registered/{subject}/{subject}_registered.nii.gz",
output_path=f"registered/{subject}/{subject}_lesion_registered_mask.nii.gz",
is_binary=True
)
# Step 2: Create probabilistic lesion atlas
template = nib.load("template.nii.gz")
lesion_maps = []
for subject in subjects:
lesion = nib.load(f"registered/{subject}/{subject}_lesion_registered_mask.nii.gz")
lesion_maps.append(lesion.get_fdata())
# Average lesion masks to create probability map
lesion_atlas = np.mean(lesion_maps, axis=0)
# Save probabilistic atlas
atlas_img = nib.Nifti1Image(lesion_atlas, template.affine)
nib.save(atlas_img, "atlas/lesion_probability_atlas.nii.gz")
print(f"Lesion atlas created from {len(subjects)} subjects")
print(f"Probability range: {lesion_atlas.min():.3f} to {lesion_atlas.max():.3f}")
Quality Control After Mask Registration
Verify mask alignment quality:
from nidataset.preprocessing import register_mask
import nibabel as nib
import numpy as np
import matplotlib.pyplot as plt
# Register mask
register_mask(
mask_path="original_mask.nii.gz",
transform_path="transforms/transformation.tfm",
registered_path="registered/scan_registered.nii.gz",
output_path="registered/mask_registered_mask.nii.gz",
is_binary=True,
debug=True
)
# Load for quality check
registered_scan = nib.load("registered/scan_registered.nii.gz").get_fdata()
registered_mask = nib.load("registered/mask_registered_mask.nii.gz").get_fdata()
# Select middle slice
mid_slice = registered_scan.shape[2] // 2
# Create overlay visualization
fig, axes = plt.subplots(1, 3, figsize=(15, 5))
# Original scan
axes[0].imshow(registered_scan[:, :, mid_slice], cmap='gray')
axes[0].set_title('Registered Scan')
axes[0].axis('off')
# Mask only
axes[1].imshow(registered_mask[:, :, mid_slice], cmap='Reds', alpha=0.8)
axes[1].set_title('Registered Mask')
axes[1].axis('off')
# Overlay
axes[2].imshow(registered_scan[:, :, mid_slice], cmap='gray')
axes[2].imshow(registered_mask[:, :, mid_slice], cmap='Reds', alpha=0.3)
axes[2].set_title('Overlay - Quality Check')
axes[2].axis('off')
plt.tight_layout()
plt.savefig('mask_registration_qc.png', dpi=150, bbox_inches='tight')
print("Quality control image saved: mask_registration_qc.png")
# Calculate mask coverage
mask_voxels = np.sum(registered_mask > 0)
total_voxels = registered_mask.size
coverage = (mask_voxels / total_voxels) * 100
print(f"\nMask Statistics:")
print(f" Total voxels: {total_voxels:,}")
print(f" Mask voxels: {mask_voxels:,}")
print(f" Coverage: {coverage:.2f}%")
Complete Preprocessing Workflow
Full pipeline from registration to mask application:
from nidataset.preprocessing import register_CTA, register_mask
import os
def complete_preprocessing_pipeline(scan_path, mask_types, output_dir):
"""
Complete preprocessing: registration and mask transformation.
Parameters
----------
scan_path : str
Path to input scan
mask_types : list
Types of masks to register (e.g., ["lesion", "vessel"])
output_dir : str
Output directory
"""
scan_id = os.path.basename(scan_path).replace('.nii.gz', '')
case_dir = f"{output_dir}/{scan_id}"
print(f"Processing {scan_id}...")
# Step 1: Register to template
print(" Registering to template...")
register_CTA(
nii_path=scan_path,
mask_path=f"masks/{scan_id}_brain_mask.nii.gz",
template_path="atlas/template.nii.gz",
template_mask_path="atlas/template_brain_mask.nii.gz",
output_path=case_dir,
cleanup=True
)
# Step 2: Register additional masks
print(" Registering additional masks...")
transform_path = f"{case_dir}/{scan_id}_transformation.tfm"
reference_path = f"{case_dir}/{scan_id}_registered.nii.gz"
registered_masks = []
for mask_type in mask_types:
mask_path = f"masks/{scan_id}_{mask_type}_mask.nii.gz"
if not os.path.exists(mask_path):
print(f" {mask_type}: not found, skipping")
continue
try:
output_path = f"{case_dir}/{scan_id}_{mask_type}_registered_mask.nii.gz"
register_mask(
mask_path=mask_path,
transform_path=transform_path,
registered_path=reference_path,
output_path=output_path,
is_binary=True
)
registered_masks.append(mask_type)
print(f" {mask_type}: registered")
except Exception as e:
print(f" {mask_type}: failed - {str(e)}")
print(f" Complete! Registered {len(registered_masks)} masks\n")
return registered_masks
# Process multiple scans
scans = ["patient001.nii.gz", "patient002.nii.gz", "patient003.nii.gz"]
mask_types = ["lesion", "vessel", "hemorrhage"]
for scan in scans:
complete_preprocessing_pipeline(
scan_path=f"raw_data/{scan}",
mask_types=mask_types,
output_dir="preprocessed"
)
Typical Workflow
from nidataset.preprocessing import register_CTA, register_mask
# 1. Register main scan
register_CTA(
nii_path="scan.nii.gz",
mask_path="scan_brain_mask.nii.gz",
template_path="template.nii.gz",
template_mask_path="template_brain_mask.nii.gz",
output_path="registered/"
)
# 2. Register additional masks using same transformation
additional_masks = [
"lesion_mask.nii.gz",
"vessel_mask.nii.gz",
"roi_mask.nii.gz"
]
for mask_file in additional_masks:
base_name = mask_file.replace("_mask.nii.gz", "")
output_name = f"{base_name}_registered_mask.nii.gz"
register_mask(
mask_path=mask_file,
transform_path="registered/scan_transformation.tfm",
registered_path="registered/scan_registered.nii.gz",
output_path=f"registered/{output_name}",
is_binary=True
)
# 3. Use registered masks for analysis
# - Voxel-wise comparisons
# - ROI statistics
# - Population studies
# - Atlas-based segmentation