generate_brain_mask
Generate a binary brain tissue mask from a medical imaging volume using intensity thresholding, morphological operations, and connected component analysis.
generate_brain_mask(
nii_path: str,
output_path: str,
threshold: tuple = None,
closing_radius: int = 3,
debug: bool = False
) -> None
Overview
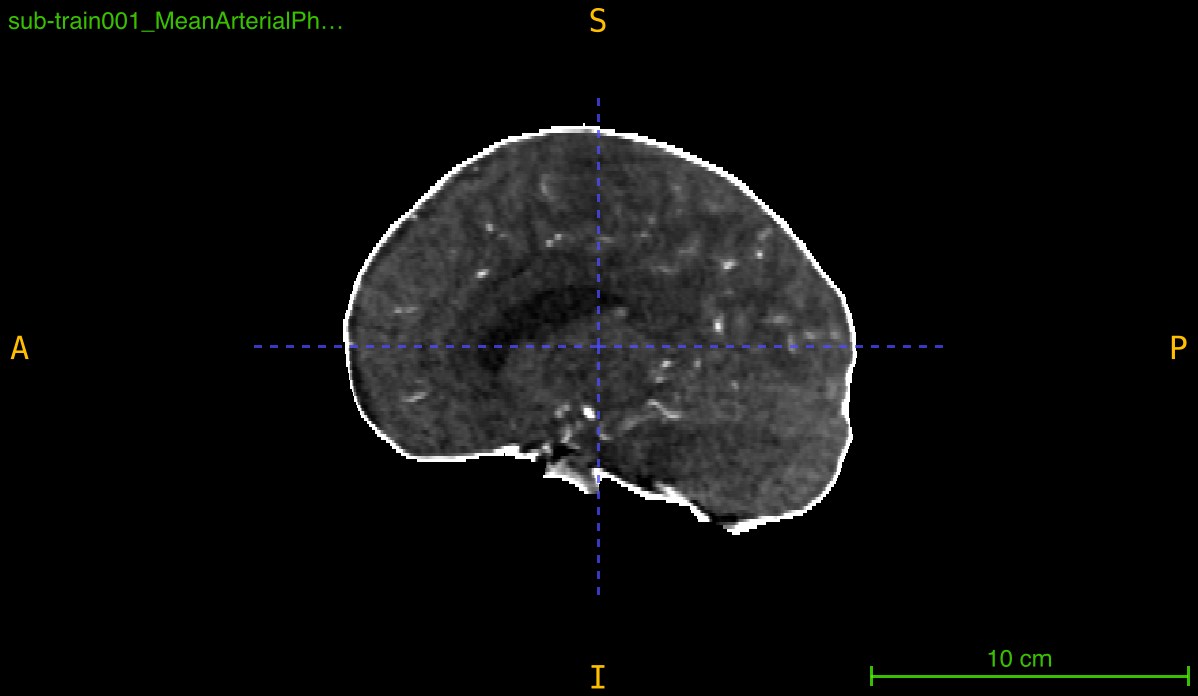
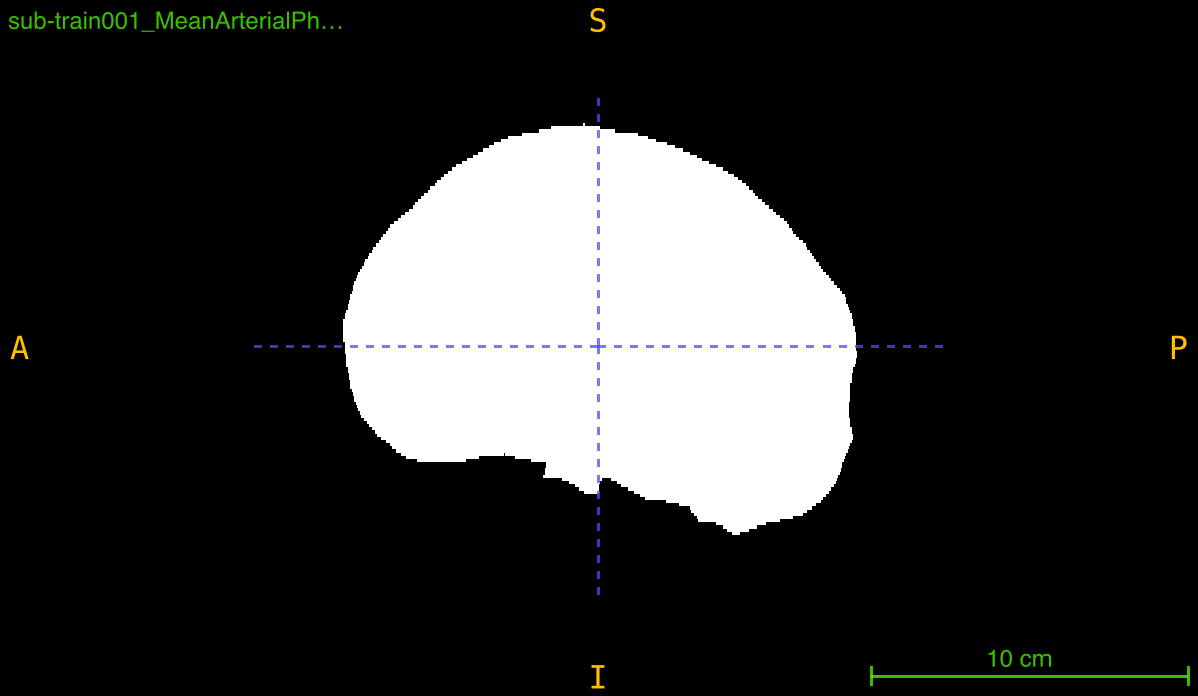
This function creates a binary mask that isolates brain tissue from background and skull by applying a multi-step segmentation pipeline:
- Thresholding: Segments tissue based on intensity values (manual or automatic Otsu)
- Hole filling: Fills small gaps within the segmented region
- Morphological closing: Smooths boundaries and connects nearby regions
- Component selection: Keeps only the largest connected component (assumed to be brain)
The result is a clean, connected binary mask suitable for skull stripping, region of interest extraction, or preprocessing pipelines.
Parameters
| Name | Type | Default | Description |
|---|---|---|---|
nii_path | str | required | Path to the input medical volume in .nii.gz format. |
output_path | str | required | Directory where the brain mask will be saved. Created automatically if it doesn’t exist. |
threshold | tuple or None | None | Intensity range (low, high) for segmentation. If None, uses automatic Otsu thresholding. |
closing_radius | int | 3 | Radius in voxels for morphological closing to refine mask boundaries. |
debug | bool | False | If True, prints diagnostic information including thresholds and file paths. |
Returns
None – The function saves the binary mask to disk.
Output File
The mask is saved as:
<PREFIX>_mask.nii.gz
where <PREFIX> is the original filename without the .nii.gz extension.
Example: Input scan_001.nii.gz → Output scan_001_mask.nii.gz
Output Mask Properties
- Data type: uint8 (8-bit unsigned integer)
- Tissue voxels: Value
1 - Background: Value
0 - Dimensions: Same as input volume
- Spatial metadata: Inherits affine transformation from input
Thresholding Strategies
Automatic Thresholding (threshold=None)
Uses Otsu’s method with adaptive bounds:
otsu_value = threshold_otsu(non_zero_voxels)
lower_bound = otsu_value × 0.5 # Relaxed to capture soft tissue
upper_bound = otsu_value × 2.0 # Extended to include bright regions
Advantages:
- Adapts to different intensity distributions
- No manual parameter tuning
- Works across different scanners and protocols
When to use: Variable intensity ranges, mixed datasets, initial exploration
Manual Thresholding (threshold=(low, high))
Uses fixed intensity bounds for all scans:
binary_mask = (intensity >= low) & (intensity <= high)
Advantages:
- Consistent segmentation across dataset
- Precise control over tissue selection
- Reproducible results
When to use: Homogeneous datasets, known intensity ranges, production pipelines
Example intensity ranges:
- CT with contrast:
(50, 300)for brain tissue - CT plain:
(20, 100)for soft tissue - MRI T1:
(50, 200)typical brain - MRI T2:
(100, 500)typical brain
Segmentation Pipeline Details
Step 1: Initial Thresholding
Creates a binary mask based on intensity values:
binary_mask = (data >= lower) & (data <= upper)
Step 2: Hole Filling
Fills internal gaps that might be caused by blood vessels or ventricles:
filled_mask = binary_fill_holes(binary_mask)
Step 3: Morphological Closing
Smooths boundaries and connects nearby regions using a spherical structuring element:
closed_mask = binary_closing(filled_mask, ball(closing_radius))
Closing radius effects:
- Radius 1-2: Minimal smoothing, preserves details
- Radius 3-4: Moderate smoothing, typical use
- Radius 5+: Aggressive smoothing, may lose small structures
Step 4: Largest Component Selection
Keeps only the largest connected component (assumed to be the brain):
labeled = label(closed_mask)
brain_mask = labeled == largest_component_label
This removes disconnected artifacts, noise, and small false positives.
Exceptions
| Exception | Condition |
|---|---|
FileNotFoundError | The input file does not exist |
ValueError | File is not in .nii.gz format |
ValueError | Input is not a 3D volume |
ValueError | Threshold tuple does not contain exactly 2 values |
Usage Notes
- Input Format: Only
.nii.gzfiles are accepted - 3D Volumes Required: Input must be a 3D NIfTI image
- Output Directory: Automatically created if it doesn’t exist
- Single Component: Only the largest connected component is retained
- Intensity Units: Ensure threshold values match your scan’s intensity scale
Examples
Basic Usage - Automatic Thresholding
Generate mask with automatic Otsu thresholding:
from nidataset.volume import generate_brain_mask
generate_brain_mask(
nii_path="scans/patient_001.nii.gz",
output_path="masks/",
threshold=None,
closing_radius=3
)
# Output: masks/patient_001_mask.nii.gz
With Debug Information
Enable verbose output to see threshold values:
generate_brain_mask(
nii_path="ct_scans/brain_scan.nii.gz",
output_path="brain_masks/",
threshold=None,
closing_radius=3,
debug=True
)
# Prints:
# Using Otsu's threshold: 125.50
# Adjusted range: (62.75, 251.00)
# Input file: 'ct_scans/brain_scan.nii.gz'
# Output path: 'brain_masks/'
# Brain mask saved at: brain_masks/brain_scan_mask.nii.gz
Manual Thresholding
Use fixed intensity bounds for consistent segmentation:
generate_brain_mask(
nii_path="data/scan.nii.gz",
output_path="data/masks/",
threshold=(50, 300), # CT brain tissue range
closing_radius=3,
debug=True
)
Fine-Tuned Morphological Processing
Adjust closing radius for different characteristics:
# Conservative closing for detailed structures
generate_brain_mask(
nii_path="high_res_scan.nii.gz",
output_path="detailed_masks/",
threshold=(40, 200),
closing_radius=2, # Minimal smoothing
debug=True
)
# Aggressive closing for noisy data
generate_brain_mask(
nii_path="noisy_scan.nii.gz",
output_path="smooth_masks/",
threshold=(50, 250),
closing_radius=6, # Strong smoothing
debug=True
)
Finding Optimal Threshold
Test different thresholds to determine best settings:
import nibabel as nib
import numpy as np
from nidataset.volume import generate_brain_mask
test_thresholds = [
(30, 200),
(50, 250),
(70, 300),
None # Automatic
]
scan_file = "test_scan.nii.gz"
for i, thresh in enumerate(test_thresholds):
output_folder = f"threshold_test/test_{i}/"
generate_brain_mask(
nii_path=scan_file,
output_path=output_folder,
threshold=thresh,
closing_radius=3,
debug=True
)
# Analyze result
mask = nib.load(f"{output_folder}/test_scan_mask.nii.gz")
mask_data = mask.get_fdata()
brain_voxels = np.sum(mask_data > 0)
total_voxels = np.prod(mask_data.shape)
coverage = (brain_voxels / total_voxels) * 100
thresh_str = f"{thresh[0]}-{thresh[1]}" if thresh else "Otsu"
print(f"Threshold {thresh_str}: Coverage = {coverage:.1f}%")
print("\nVisually inspect outputs to select best threshold")
Quality Control Verification
Generate mask and verify quality:
import nibabel as nib
import numpy as np
from nidataset.volume import generate_brain_mask
# Generate mask
generate_brain_mask(
nii_path="qa/test_scan.nii.gz",
output_path="qa/masks/",
threshold=(50, 300),
closing_radius=3,
debug=True
)
# Load and verify
scan = nib.load("qa/test_scan.nii.gz")
mask = nib.load("qa/masks/test_scan_mask.nii.gz")
scan_data = scan.get_fdata()
mask_data = mask.get_fdata()
# Quality metrics
brain_voxels = np.sum(mask_data > 0)
total_voxels = np.prod(mask_data.shape)
coverage = (brain_voxels / total_voxels) * 100
# Check mask properties
unique_values = np.unique(mask_data)
num_components = len(np.unique(mask_data)) - 1 # Exclude background
print(f"\nQuality Control Report:")
print(f" Scan shape: {scan_data.shape}")
print(f" Mask shape: {mask_data.shape}")
print(f" Brain coverage: {coverage:.1f}%")
print(f" Mask values: {unique_values}")
print(f" Number of components: {num_components}")
# Expected values
if coverage < 20 or coverage > 60:
print(" Warning: Unusual coverage, review threshold")
if num_components > 1:
print(" Warning: Multiple components detected")
else:
print(" Mask looks good")
Applying Mask to Original Scan
Use mask for skull stripping:
import nibabel as nib
from nidataset.volume import generate_brain_mask
scan_file = "scans/patient_042.nii.gz"
mask_output = "masks/"
stripped_output = "skull_stripped/"
# Generate mask
generate_brain_mask(
nii_path=scan_file,
output_path=mask_output,
threshold=(50, 300),
closing_radius=4,
debug=True
)
# Load scan and mask
scan = nib.load(scan_file)
scan_data = scan.get_fdata()
mask = nib.load(f"{mask_output}/patient_042_mask.nii.gz")
mask_data = mask.get_fdata()
# Apply mask (skull stripping)
stripped_data = scan_data * mask_data
# Save skull-stripped scan
os.makedirs(stripped_output, exist_ok=True)
stripped_img = nib.Nifti1Image(stripped_data, scan.affine)
nib.save(stripped_img, f"{stripped_output}/patient_042_stripped.nii.gz")
print("Skull stripping complete")
Comparing Different Closing Radii
Test morphological processing parameters:
import nibabel as nib
import numpy as np
from nidataset.volume import generate_brain_mask
radii = [2, 3, 4, 5, 6]
scan_file = "test_data/scan.nii.gz"
for radius in radii:
output_folder = f"closing_test/radius_{radius}/"
generate_brain_mask(
nii_path=scan_file,
output_path=output_folder,
threshold=(50, 300),
closing_radius=radius,
debug=True
)
# Analyze smoothness (via surface area proxy)
mask = nib.load(f"{output_folder}/scan_mask.nii.gz")
mask_data = mask.get_fdata()
# Count edge voxels
from scipy import ndimage
edges = ndimage.sobel(mask_data)
edge_voxels = np.sum(edges > 0)
print(f"Radius {radius}: Edge voxels = {edge_voxels}")
print("\nSmaller edge count = smoother mask")
Integration with Preprocessing Pipeline
Use masks in a complete preprocessing workflow:
import nibabel as nib
import numpy as np
from nidataset.volume import generate_brain_mask
def preprocess_scan(input_path, output_path):
"""Complete preprocessing with skull stripping and normalization."""
# Step 1: Generate brain mask
generate_brain_mask(
nii_path=input_path,
output_path="temp_masks/",
threshold=(50, 300),
closing_radius=3,
debug=True
)
# Step 2: Load scan and mask
scan = nib.load(input_path)
scan_data = scan.get_fdata()
filename = os.path.basename(input_path)
mask_file = filename.replace('.nii.gz', '_mask.nii.gz')
mask = nib.load(f"temp_masks/{mask_file}")
mask_data = mask.get_fdata()
# Step 3: Apply mask
brain_only = scan_data * mask_data
# Step 4: Normalize intensities within mask
brain_voxels = brain_only[mask_data > 0]
mean_intensity = brain_voxels.mean()
std_intensity = brain_voxels.std()
normalized = np.zeros_like(brain_only)
normalized[mask_data > 0] = (brain_voxels - mean_intensity) / std_intensity
# Step 5: Save preprocessed scan
preprocessed = nib.Nifti1Image(normalized, scan.affine)
nib.save(preprocessed, output_path)
print(f"Preprocessed scan saved to {output_path}")
return output_path
# Use in pipeline
preprocess_scan(
"raw_scans/patient_001.nii.gz",
"preprocessed/patient_001_preprocessed.nii.gz"
)
Batch Processing with Parameter Optimization
Process dataset with optimized parameters per scan type:
from nidataset.volume import generate_brain_mask
import json
# Load scan metadata
with open('scan_metadata.json', 'r') as f:
metadata = json.load(f)
for scan_id, info in metadata.items():
scan_file = f"scans/{scan_id}.nii.gz"
# Select parameters based on scan type
if info['modality'] == 'CT_contrast':
thresh = (50, 300)
radius = 3
elif info['modality'] == 'CT_plain':
thresh = (20, 100)
radius = 4
elif info['modality'] == 'MRI_T1':
thresh = None # Automatic
radius = 3
else:
continue
generate_brain_mask(
nii_path=scan_file,
output_path=f"masks/{info['modality']}/",
threshold=thresh,
closing_radius=radius,
debug=True
)
print(f"Processed {scan_id} ({info['modality']})")
Creating Visualization Overlay
Generate mask overlay for quality inspection:
import nibabel as nib
import numpy as np
from nidataset.volume import generate_brain_mask
scan_file = "visualization/scan.nii.gz"
# Generate mask
generate_brain_mask(
nii_path=scan_file,
output_path="visualization/masks/",
threshold=(50, 300),
closing_radius=3
)
# Load scan and mask
scan = nib.load(scan_file)
scan_data = scan.get_fdata()
mask = nib.load("visualization/masks/scan_mask.nii.gz")
mask_data = mask.get_fdata()
# Create overlay with mask boundary highlighted
from scipy import ndimage
edges = ndimage.sobel(mask_data)
overlay = scan_data.copy()
overlay[edges > 0] = scan_data.max() # Bright edge highlighting
# Save overlay
overlay_img = nib.Nifti1Image(overlay, scan.affine)
nib.save(overlay_img, "visualization/overlay_with_mask.nii.gz")
print("Overlay created for visual inspection in viewer")
Typical Workflow
from nidataset.volume import generate_brain_mask
import nibabel as nib
import numpy as np
# 1. Generate brain mask
scan_file = "data/patient_scan.nii.gz"
mask_output = "data/masks/"
generate_brain_mask(
nii_path=scan_file,
output_path=mask_output,
threshold=(50, 300), # Adjust for your modality
closing_radius=3,
debug=True
)
# 2. Verify mask quality
mask = nib.load("data/masks/patient_scan_mask.nii.gz")
mask_data = mask.get_fdata()
coverage = np.sum(mask_data > 0) / np.prod(mask_data.shape) * 100
print(f"Brain coverage: {coverage:.1f}%")
# 3. Use mask for:
# - Skull stripping
# - Region of interest extraction
# - Focused analysis on brain tissue
# - Preprocessing before segmentation or classification